Chapter 11: Lessons from Adjacent Industries — AI in Pharma, Healthcare, and Nutraceuticals
Why this chapter
Cosmetics is the small player in microbiome industrialization. Sequencing, generative protein design, in silico trial simulation, defined consortia — every tool we covered in Part II and Part III — was not built for cosmetics. Adjacent industries with larger capital, addressing harder problems (IPF, recurrent C. difficile infection, diabetes, cancer), built them first — and cosmetics transfers them in. That is not a weakness — it is the cheapest R&D strategy available. A single round of AI drug discovery costs near the annual R&D budget of a global cosmetic major; letting someone else build the tooling and porting it later is therefore rational.
Brief B explicitly asks this chapter — "tell me how adjacent industries industrialized microbiome × AI." The honest answer is that pharma and nutraceuticals carry operational templates cosmetics has not yet replicated, in five categories: (i) end-to-end clinical readout for an AI-designed active (Insilico / rentosertib), (ii) FDA approval for a microbiome therapeutic (Seres / VOWST), (iii) clinical trial of a defined-consortium product (Vedanta / VE303), (iv) personalization built on a microbiome × phenotype data cohort (ZOE PREDICT), (v) industry adoption of generative protein design SaaS (Cradle, Profluent). None transfer to cosmetics one-to-one — naming what does and does not transfer is this chapter's work.
Three quantitative anchors for this chapter 1. End-to-end AI drug discovery timeline: Insilico Medicine's rentosertib went from AI target identification to IND in 30 months, to Phase 2a in ~5 years [25]. The 12-week Phase 2a reported a +98.4 mL FVC change at 60 mg QD vs −20.3 mL on placebo [14]. Cosmetic-industry equivalent publications: 0 (Gap 1). 2. FDA-approved microbiome therapeutics: as of May 2026, two — Seres's VOWST (SER-109, approved April 2023) [7] and Ferring's Rebyota (RBX2660, approved November 2022). Cosmetic equivalents: 0 — though Geltor's PrimaColl received the first GRAS "no questions" letter for a biodesigned cosmetic-adjacent ingredient in October 2025 on the ingestible side [8] (Gap 5). 3. Adjacent transfer asymmetry: pharma majors (Pfizer, Novo Nordisk, GSK) disclose AI partnerships but no microbiome-headlined programs (Gap 11). Inversely, microbiome-headlined companies (Seres, Vedanta) are mid-caps, not majors. Cosmetic R&D planners can borrow the AI partnership pattern and the microbiome regulatory pattern — but no single big company combines both.
11.1 Framing — why adjacent industries are cosmetics' cheapest R&D
Three asymmetries position cosmetics on the import side.
Capital asymmetry. A single small-molecule drug discovery program is traditionally estimated at $1–2B; AI-accelerated pipelines compress that by half or more, but the residual is still on the order of a global cosmetic major's entire annual R&D budget. L'Oréal's group R&I budget was ~€1.27B in 2024; Amorepacific's was ~₩280B (~$200M). The cosmetic industry's capital structure cannot afford to train an AlphaFold-class model de novo, build a million-data-point ChEMBL-class corpus, or run an Insilico-style end-to-end pipeline — that infrastructure must come from elsewhere.
Regulatory asymmetry. FDA, EMA, and Korea MFDS already have an evaluation framework for microbiome drugs — incomplete but extant. Seres's VOWST is the first case showing how FDA evaluates a live-microbiome product's efficacy, CMC, and environmental risk [27]. Cosmetic regulators have no equivalent framework (Gap 5; Chapter 7). If cosmetics moves toward live engineered strains, it will have to cite pharma precedent because no direct path exists — that, too, is adjacent-industry dependency.
Data asymmetry. ChEMBL (2.4M assays), PubChem (110M+ compounds), PDB (220K structures) are assets pharma released as public goods over a century. Cosmetics has no equivalent (Gap 12). It is rational for cosmetic AI to fine-tune from pharma public data and to do domain adaptation where the two industries' metabolite spaces partially overlap — because cosmetics has no capital to generate that data from scratch.
The three asymmetries make adjacent-industry dependence a permanent structural condition, not a stopgap. Chapter 12's blueprint assumes this — the question a cosmetic R&D planner asks is not "how do we build it ourselves?" but "which tool do we borrow from where, and what can we not borrow?"
11.2 Insilico Medicine — the end-to-end clinical template for AI drug discovery
The single data point cosmetics should most envy from adjacent industries is [25]'s Nature Biotechnology paper. Insilico Medicine ran PandaOmics (target-identification AI) and Chemistry42 (generative molecule-design AI) in series to identify TNIK kinase as a novel target for idiopathic pulmonary fibrosis (IPF), and to design a first-in-class small-molecule inhibitor, rentosertib (ISM001-055), for that target. From target hypothesis to IND submission: 18 months. From IND to Phase 1 first-in-human: another 12 months. Total: 30 months — one-half to one-quarter of the traditional target-to-IND timeline of 4–6 years [25].
One year later, [14]'s Nature Medicine Phase 2a results provided the first peer-reviewed answer to whether an AI-discovered molecule produces measurable effects in clinical patients. The randomized double-blind placebo-controlled trial enrolled 71 IPF patients with 12 weeks of dosing. Primary endpoint: forced vital capacity (FVC) change. Result — the rentosertib 60 mg QD arm showed +98.4 mL FVC vs −20.3 mL on placebo, significant even on top of standard of care (nintedanib, pirfenidone). Safety profile was acceptable; two patient deaths were adjudicated within the natural-history range of IPF. This is the first peer-reviewed case of an end-to-end AI-discovered molecule reaching a clinical efficacy signal.
Transferability to cosmetics is structurally clear and operationally empty. Structure: (i) AI-accelerated target identification — microbiome metabolite or host protein (Chapter 4); (ii) generative active-molecule design — the AlphaFold/Boltz/Chai stack or its ESM3 equivalent (Chapter 5); (iii) in-house wet-lab validation (Chapter 7 DBTL); (iv) ex vivo / clinical readout (Chapter 9). The tooling for every step became commodity in 2024. What is empty is that the cosmetic industry has not run the full pipeline through to publication (Gap 1). [11]'s PRISMA review names this explicitly — zero of 74 included studies report a published clinical readout of an AI-designed cosmetic active.
It is worth noting that [33]'s GENTRL DDR1 paper opened this arc. GENTRL was a reinforcement-learning generative model that designed, synthesized, and in-vitro tested six DDR1 inhibitors in 21 days — Nature Biotechnology. Six years later, the same company did the same thing for IPF patients — that timeline calibrates what cosmetics can borrow. Cosmetic adoption of AI tooling is stuck near the first method-paper stage — the 2021 [4] IBM-Unilever paper, equivalent in conceptual maturity to GENTRL in 2019. The clinical-readout step is missing.
Two specifics a cosmetic R&D planner can take from Insilico:
First, Insilico's publication strategy. They published methodology at every stage — the PandaOmics and Chemistry42 algorithm papers, the Nature Biotechnology clinical-entry paper, the Nature Medicine Phase 2a paper. IP protection was concentrated on the molecule itself and synthesis routes; methodology was open. Cosmetic majors follow the opposite pattern — both methodology and molecules stay in-house. The publication asymmetry means cosmetic AI ends up not externally evaluated (Gap 15).
Second, the organizational meaning of end-to-end. Insilico's AI team, synthesis team, and clinical team are a single organizational unit. Cosmetic majors typically separate the AI team in R&I from the clinical (or efficacy-testing) team. What made the 30-month timeline possible was not the algorithm — it was organizational integration. That is the structural opportunity for Korean OEMs (COSMAX, Kolmar) — B2B-internal clinical infrastructure already exists; bolting AI on top is feasible.
11.3 Recursion, Exscientia, and the AI drug-discovery SaaS landscape
Beyond Insilico, two companies have driven AI drug discovery industrialization with different models. Recursion Pharmaceuticals is a phenomics company — automated cellular-image arrays measure phenotypic profiles of compounds, and ML is trained on that data. The [24] merger (announced August 2024, completed November 2024) integrated phenomics (Recursion) with generative chemistry (Exscientia, UK-based) into a single platform. The Roche partnership focuses on neuro indications; the combined entity holds 30+ active programs.
Cosmetic transferability: phenomics infrastructure is indirectly useful. Standard in-vitro endpoints for cosmetic-active screening (fibroblast proliferation, collagen synthesis, MMP inhibition, antioxidant capacity) map naturally to Recursion-style automated imaging. Adoption in CDMO/OEM channels would partially route around the capital asymmetry — a phenotype model Recursion already trained, fine-tuned to cosmetic endpoints. The Exscientia generative-chemistry side overlaps more directly with the cosmetic-metabolite design discussion in Chapters 4 and 5.
Cradle Bio (Zurich/Delft-based) is a more cosmetic-friendly SaaS model. ML-driven protein design — for enzymes, antibodies, and biologics — accessed by customer firms through a web interface. [6]'s $73M Series B (November 2024) named [22] as a disclosed customer. The exact protein Novo is engineering with Cradle is not disclosed — GLP-1-class peptide engineering is the most plausible candidate. Borrowing the same SaaS for cosmetic R&D is cost-rational: vegan collagen (the class containing Chapter 7's Geltor PrimaColl), engineered peptides (elastin mimetics, MMP-inhibitor peptides), and microbiome-metabolite biosynthetic enzymes can be designed externally — proteins for which the cosmetic industry has no capital to build a training corpus directly.
Profluent (Berkeley, US) is the more ambitious variant. [26]'s OpenCRISPR-1 is a de novo-generated Cas9-equivalent protein — the ProGen2 protein language model, trained on the CRISPR-Cas universe, generated a synthetic protein that retains in-vitro CRISPR activity. Cosmetic near-term application is limited, but the longer-horizon implication is large: if biosynthetic enzymes for microbiome metabolite pathways can be generated the same way, the design step of Chapter 7's DBTL loop becomes commodity. Cosmetics will never build that model directly, but it is plausible to rent it via SaaS within five years.
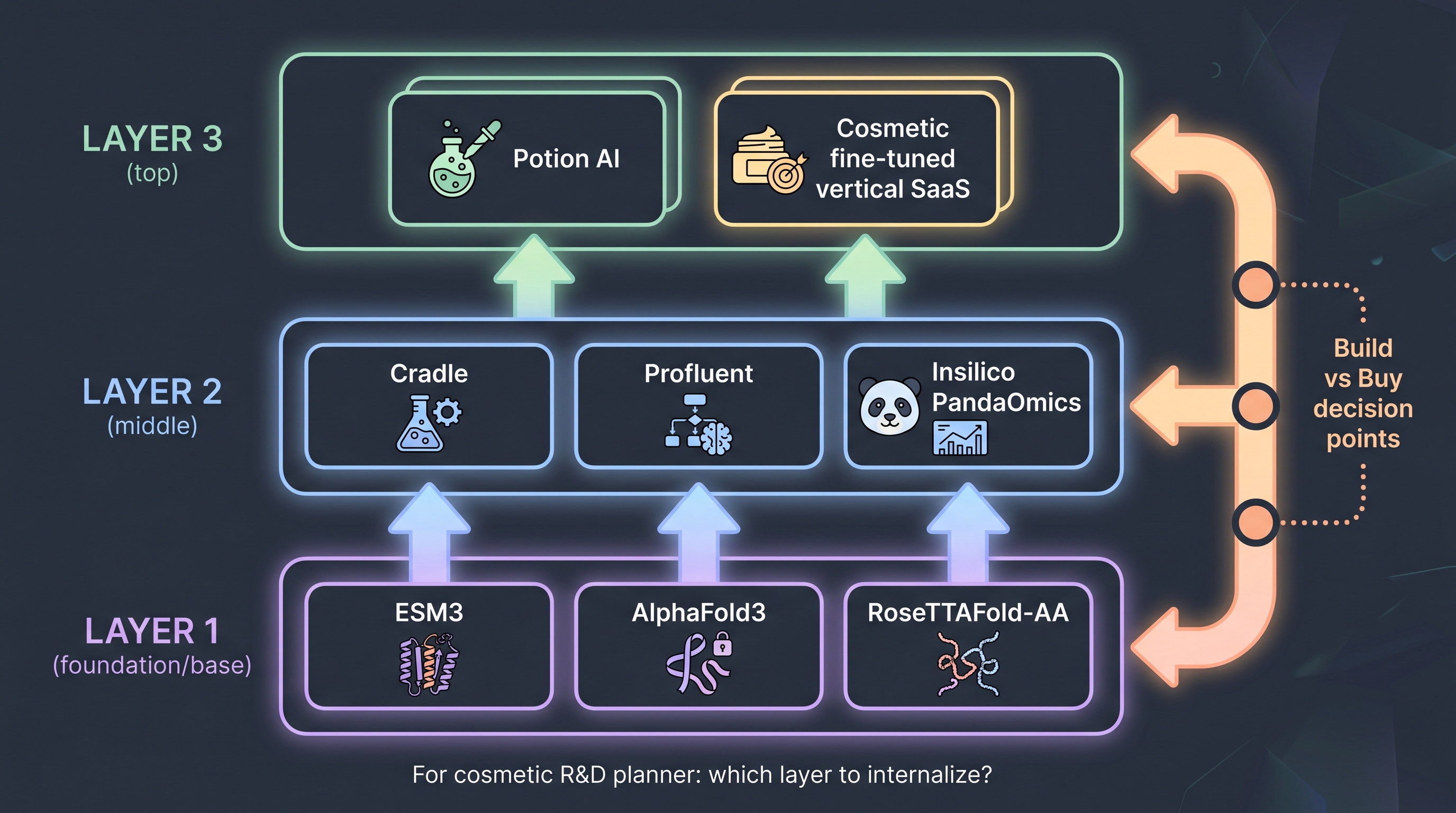
[12]'s ESM3 — a 98B-parameter protein language model with a 7B-parameter open release — is the foundation-model layer of this SaaS landscape. Released by EvolutionaryScale, ESM3 simulates 500 million years of evolution to generate de novo proteins; application companies like Cradle and Profluent fine-tune on top. The decision a cosmetic R&D planner faces is which layer to build in-house and which layer to rent. Foundation models: rent (capital and data missing). Application layer (cosmetic-endpoint-specific fine-tunes): partially in-house (in-house phenotype data is IP). UI/workflow layer: rent (Potion AI-style) or build (a choice).
This layer separation reappears as the central decision variable in the transfer-learning matrix (§11.8).
11.4 Seres VOWST — the first FDA-approved microbiome therapeutic
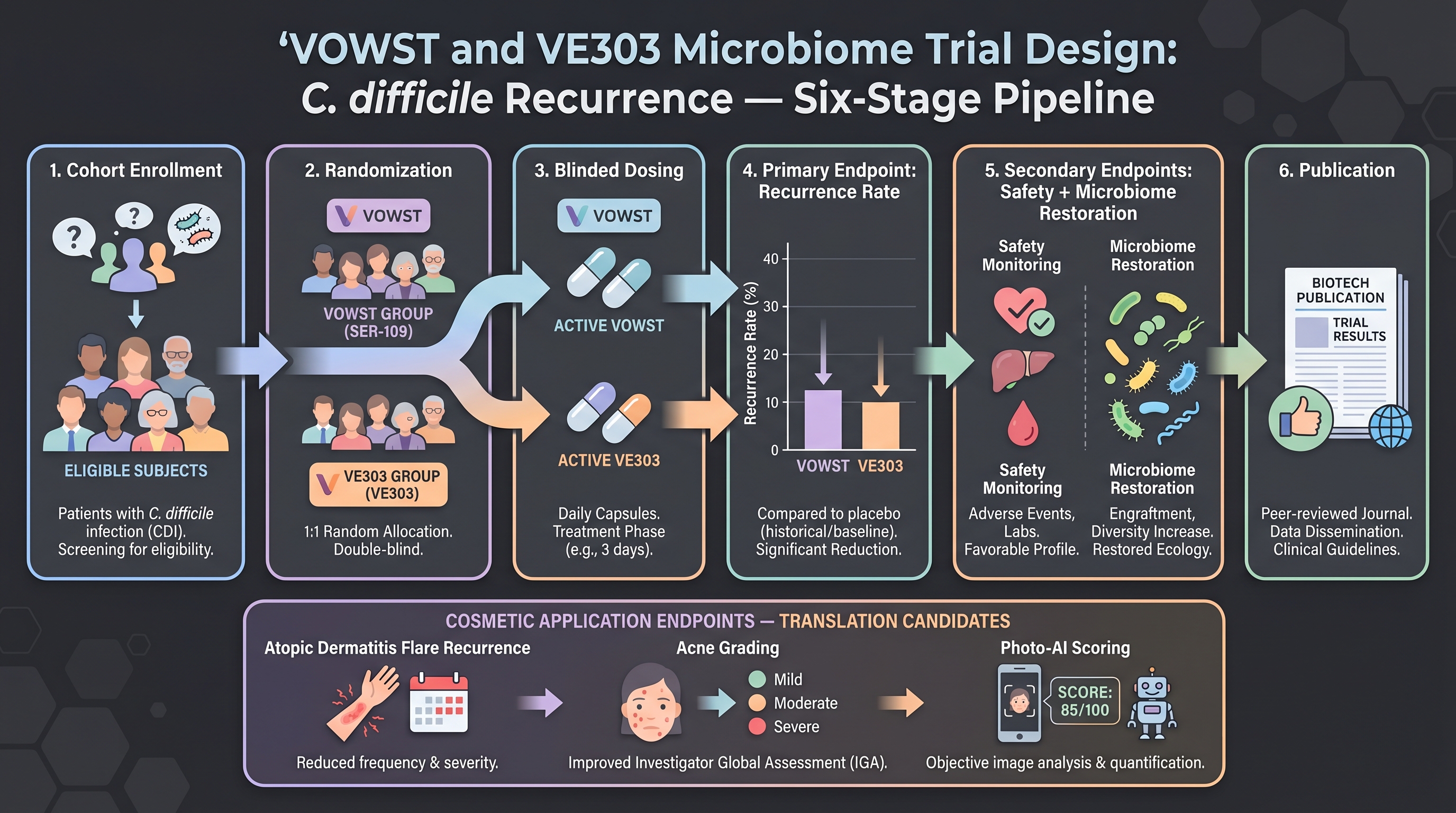
If cosmetics asks what the regulatory path looks like for a live-microbiome product, the single most important reference is Seres Therapeutics's SER-109 / VOWST. SER-109 is a purified Firmicutes-spore preparation that restores a patient's microbiome to prevent recurrence of Clostridioides difficile infection (rCDI).
The pivotal data are in [7] NEJM — the ECOSPOR III randomized double-blind placebo-controlled trial. 182 patients, 1:1 randomization. rCDI recurrence at 8 weeks — SER-109 12% vs placebo 40% (relative risk 0.32, 95% CI 0.18–0.58, P < 0.001). Number needed to treat ~4 in a single trial. That is an effect size of a different order than any cosmetic active has reported (because rCDI is a medical-emergency endpoint, not because AI is involved — the Seres pipeline is not AI-heavy). In April 2023 the FDA approved VOWST as the first oral microbiome therapeutic, and a Nestlé Health Science commercial partnership launched shortly after [27].
Cosmetic transferability: not direct. SER-109's mechanism is indirect microbiome restoration — purified spores fill the patient's missing community niche to prevent C. difficile recolonization. The closest cosmetic analog is live-commensal supplementation, which is a category cosmetic regulation does not currently accommodate (Gap 5). VOWST was approved as a drug — it followed FDA's Biologics License Application (BLA) path.
But the trial design itself transfers directly to cosmetics. Specifics:
Endpoint selection. Primary endpoint set as a single binary outcome (recurrence), unambiguously. Cosmetic trials suffer endpoint blur — "wrinkle + hydration + radiance all improved"-type vague composites. Pre-registering a single objective binary (or quantitative continuous) endpoint, ECOSPOR-III-style, is the fastest path for cosmetic efficacy work to reach peer review.
CMC (chemistry, manufacturing, controls) standards. Batch-to-batch consistency, purity, and potency measurement for live-strain products is a trail Seres blazed with FDA. Strain identity (genome sequencing), viable count, and contamination panel are all standardized. If cosmetics moves toward live-strain products (or strongly strain-defined postbiotics), Seres's CMC dossier is the reference template.
Cohort design. ECOSPOR III used multi-site enrollment to avoid over-fitting to a single ethnicity or geography. This pre-empts the most frequent criticism of cosmetic trials — non-representative Fitzpatrick types (Gap 4).
Vedanta Biosciences's VE303 is the second attempt in the same category, with a different molecular-biology approach. [17] in JAMA reports VE303's Phase 2 CONSORTIUM trial — at 8 weeks, high-dose VE303 showed an odds ratio of 0.27 (95% CI 0.09–0.81) for rCDI recurrence vs placebo. Unlike SER-109's donor-derived spore preparation, VE303 is a defined bacterial consortium — eight Clostridia strains — which is more amenable to industrial standardization. In May 2024, [29] announced first-patient-dosed for the Phase 3 RESTORATiVE303 trial.
Vedanta shows the industrial justification for defined consortia — a path to batch-to-batch consistency without donor variability. For a cosmetic R&D planner designing a microbiome cocktail product, the right reference is not a donor-derived or fermentate-derived single strain — it is the VE303 model (defined N-strain consortium, every strain genome-sequenced, ratios quantified). Chapter 7's DBTL loop makes such consortium design directly feasible.
A cautionary tale to keep visible: Vedanta's VE202 (inflammatory bowel disease target) missed its Phase 2 ulcerative colitis primary endpoint in 2022 — a signal that the defined-consortium approach does not work for every indication. When cosmetics applies the same model, the precondition is that the clinical endpoint must be mechanistically sensitive to microbiome manipulation. rCDI is an endpoint where microbiome deficit is the causal driver — that is why SER-109 and VE303 work. In cosmetics, equivalent "microbiome-as-causal-driver" endpoints are likely a subset of atopic dermatitis and acne — and not general anti-aging (Gap 14).
[18]'s Experimental Dermatology review provides the academic-grade landscape of microbiome therapeutics for dermatological indications. The clearest clinical-readout grade is atopic dermatitis × Roseomonas mucosa / Staphylococcus hominis A9 — the Gallo lab's [20]. That is the closest peer-reviewed "live-microbe cosmetics" precedent and ties into Chapter 9's ground-truth line.
11.5 Pharma majors and microbiome — the absence of disclosure
Brief B asks how pharma majors (Pfizer, Novo Nordisk, Eli Lilly) industrialize microbiome × AI. The honest answer is that they do not disclose microbiome-headlined programs (Gap 11). What they disclose is AI partnerships — and the exact indication and molecule those partnerships target is opaque.
Pfizer × XtalPi. [23]'s June 2025 expanded partnership announcement extends a collaboration begun ~8 years earlier. XtalPi is a China/US-based AI drug-discovery + materials-science simulation platform offering crystal structure prediction, drug-target interaction modeling, and active-molecule design. Pfizer's disclosed use cases are general small-molecule discovery; no microbiome-specific program is disclosed. (Pfizer separately had an earlier microbiome partnership with Second Genome that has been quiet since early 2020.)
Novo Nordisk × Cradle Bio. [22] was named as a Cradle customer in the Series B disclosure. GLP-1 peptide engineering (semaglutide/tirzepatide-class) is the plausible use case, but it is not disclosed. Novo runs no public microbiome-specific program — however, the GLP-1-microbiome interaction (SCFA-mediated GLP-1 secretion) is well-described in the academic literature, providing indirect evidence that an in-house program exists.
Eli Lilly. By public disclosure, no microbiome-specific AI program exists. Lilly's AI strategy — including the 2024 Isomorphic Labs (Alphabet) partnership — centers on protein design and small-molecule discovery.
GSK × Fleming Initiative. [9]'s £45M Imperial College-based program targets antimicrobial resistance (AMR) with AI. AMR is microbiome-adjacent but not a cosmetic-direct use case — though predicting antibiotic-resistant strains is mechanistically related to the cosmetic-preservative-resistance problem.
Roche / Recursion. After the 2024 merger, Recursion-Exscientia retains the Roche partnership but emphasizes neuroscience and oncology over microbiome-adjacent indications.
What cosmetic R&D planners should take from this landscape: pharma majors' microbiome-headline programs cannot be borrowed because they do not exist. What can be borrowed is the AI partnership pattern. Pfizer-XtalPi, Novo-Cradle, GSK-Fleming all share the same structure — internal R&D core + external AI SaaS partner. Cosmetic majors (L'Oréal-Microsoft, Shiseido-Accenture) have already adopted that pattern — but disclosed microbiome-specific use cases remain few.
[14] — by virtue of being publicly traded (HKEX IPO March 2025) — is the most auditable case study. The Insilico 2024 IPO prospectus disclosed 18 clinical programs, 5 in Phase 2, 100+ AI scientists, and $300M+ cumulative R&D spend. Cosmetic majors' disclosures — kept internal — are not comparable. If the cosmetic industry wants a publicly auditable comparable, a Korean OEM like COSMAX (whose B2B model creates publishing incentive) is the most plausible candidate.
11.6 ZOE PREDICT — the template for personalized nutrition and cosmetic personalization
Brief B asks for nutraceuticals explicitly. In that category, the single most important reference series is ZOE's PREDICT cohort.
[2] Nature Medicine — PREDICT 1, 1,098 deeply phenotyped individuals — showed personalized relationships between microbiome composition and host metabolic response (post-meal glycemic and lipemic response). Subsequently, [3] reported cohort updates, [31] Nature Medicine reported the personalized-nutrition intervention (METHOD trial), and [32] Nature reported microbiome health-ranking findings (centering Faecalibacterium prausnitzii, Roseburia, others).
Key components of the ZOE model:
(i) Consumer-grade home test kit. Stool sample + dried blood spot + continuous glucose monitor patch. Users collect at home under a standardized protocol and return by mail.
(ii) ML personalization engine. Microbiome × diet × glucose timeline → individualized dietary recommendations. The algorithm was validated to clinical grade via RCT [31].
(iii) Ongoing cohort operation. Operates at 100,000+ scale; models retrain over time; new microbe-phenotype associations reach Nature-grade publication.
Transfer to cosmetic personalization: this is the reference template for Chapter 6's digital twin and Chapter 8's POND'S in-store diagnostic. The in-store diagnostic mimics ZOE's (i); the AI recommender mimics (ii); but the ongoing-cohort learning of (iii) is not yet present in cosmetics. POND'S and COSMAX × HelloBiome [5] have adopted (i) and (ii) but lag ZOE in cohort operation scale and peer-reviewed publication cadence.
Two specifics ZOE teaches cosmetics directly:
First, the data + algorithm + RCT trio. ZOE did not launch with just a consumer test kit. The credibility moat came from building academic-PI-led (Tim Spector, Nicola Segata) cohort data and peer-reviewed publications in tandem. A cosmetic personalization startup chasing ZOE-grade trust needs the same trio (a partial fix for the audit absence in Gap 15).
Second, the phenotype asymmetry of capturing diet timing and habits. Cosmetic personalization typically captures microbiome × skin phenotype only. Systemic phenotype — diet, sleep, stress — is usually not captured. ZOE captures systemic phenotype and combines it with microbiome — that is why ZOE's personalization signal-to-noise is stronger. If cosmetic R&D — particularly in the routine-heavy K-beauty user base — captured the same systemic phenotype, ZOE-equivalent personalization quality would become possible (a Chapter 12 blueprint thread).
Viome and DayTwo are adjacent variants. Viome markets metatranscriptomics (measuring active function) as a differentiator, but its peer-reviewed cohort publication cadence is weaker than ZOE's. DayTwo spun out of the Weizmann Institute's Cell paper on Personalized Nutrition by Prediction of Glycemic Responses (Zeevi et al. 2015) and adopted a B2B (insurer/healthcare provider) model.
If the cosmetic personalization market matures formally in the 2027–2030 window, the marketing, regulatory-arbitrage, and cohort-operations patterns these three companies learned in consumer-facing non-cosmetic microbiome personalization will likely transfer to cosmetics as-is.
11.7 Microbiome data infrastructure — public goods adjacent industries built
Three adjacent-industry public goods are worth naming because cosmetics can use them immediately, at no cost.
MGnify (operated by EMBL-EBI). A unified analysis platform for >700,000 metagenomic samples. Supports metabarcoding, shotgun metagenomics, and metatranscriptomics. Cosmetic R&D can upload in-house 16S/shotgun data to MGnify for standardized taxonomic and functional profiling, or compare against published reference cohorts (EMP, HMP). It already appeared in Chapter 3 when discussing metagenomic pipelines — but its public-data justification analogous to ZOE-grade openness is underused by cosmetics.
MIBiG (Minimum Information about a Biosynthetic Gene cluster). A curated database of >2,500 natural-product biosynthetic gene clusters. Appeared in Chapter 4 as a training corpus for metabolite screening. When the cosmetic industry trains a BGC (biosynthetic gene cluster) prediction model (antiSMASH, etc.), MIBiG is the direct ground truth. Built as a public good by academia; cosmetic R&D can simply copy the pharma + industrial-biotech usage pattern.
NPAtlas (Natural Products Atlas). A curated database of >30,000 natural product structures. When protein-ligand models like ESM3 or Chai-1 are fine-tuned for cosmetic-metabolite × host-protein docking prediction, NPAtlas is the reference for chemical space coverage. Non-profit, academia-operated.
All three databases are built and kept free by pharma + academia. Under the capital asymmetry of §11.1, the most rational first move for cosmetics is to use this infrastructure rather than build its own. But no cosmetic-specific equivalent exists (Gap 12) — no curated database of cosmetic ingredient × microbiome interaction, no benchmark of cosmetic-active × in-vitro endpoint. Chapter 12's blueprint frames this gap as an explicit opportunity vector.
11.8 What does not transfer — asymmetries of regulation, capital, and timeline
If the chapter does not name what does not transfer, it over-promises. Three non-transferable items:
Regulatory rigor asymmetry. FDA IND, Phase 1/2/3, NDA, and BLA processes are evaluation standards the cosmetic industry cannot replicate — neither the capital structure, the timeline, nor the incentive. A single SER-109 ECOSPOR III trial costs orders of magnitude more than a typical cosmetic efficacy trial. Cosmetic R&D can borrow SER-109's methodology (trial design, CMC) — but not its evaluative rigor. AI acceleration narrows this gap only partially; it does not close the capital gap.
Capital-intensity asymmetry. Cumulative R&D spend on Insilico's single molecule (rentosertib) is disclosed at $50–100M. That order of magnitude is comparable to a cosmetic company's entire annual active-ingredient portfolio R&D spend. Cosmetics cannot spend Insilico's single-molecule capital on a single active. The viable path is a portfolio model — spreading one unit of capital across many actives concurrently — which combines naturally with renting the SaaS layer (Cradle, Profluent).
Timeline asymmetry. Insilico's 2014 founding → 2018 first lead → 2024 Phase 2a → 2025 Nature Medicine is an 11-year timeline. Cosmetics has faster market cycles (2–3-year SKU lifecycle) and lacks the patient capital to sustain an 11-year R&D arc. Again, the portfolio model is the answer. Insilico's single-molecule deep dive does not transfer directly.
Intrinsic difference in clinical endpoints. rCDI (SER-109), IPF (rentosertib), and diabetes prevention (ZOE) are all medical-emergency or near-emergency endpoints. Effect sizes are large, the regulatory bar is high, and the payor is insurance. Cosmetic endpoints (wrinkle, hydration, radiance) are consumer-preference endpoints — effect sizes are small, the regulatory bar is low, the payor is the consumer. The same AI acceleration produces different outcomes against signals that differ by orders of magnitude. If cosmetic R&D planners do not model this difference explicitly, they will overclaim Insilico-style 30-month timelines for cosmetics.
Finally, [Galderma] — the largest pure-play global dermatology firm, owner of Cetaphil — deserves naming for its absence of disclosure (Gap 10). Since the March 2024 SIX (Zurich) IPO, Galderma has not publicly disclosed AI / microbiome programs. Two interpretations are possible — (i) genuine non-investment, (ii) IR-conservative posture post-IPO suppressing disclosure. In either case — for a cosmetic R&D planner wanting to reference a global dermatology specialist's AI-microbiome bets — no such reference is currently available. That itself is market signal: even the nearest medical-cosmetic adjacent firm has not bet publicly.
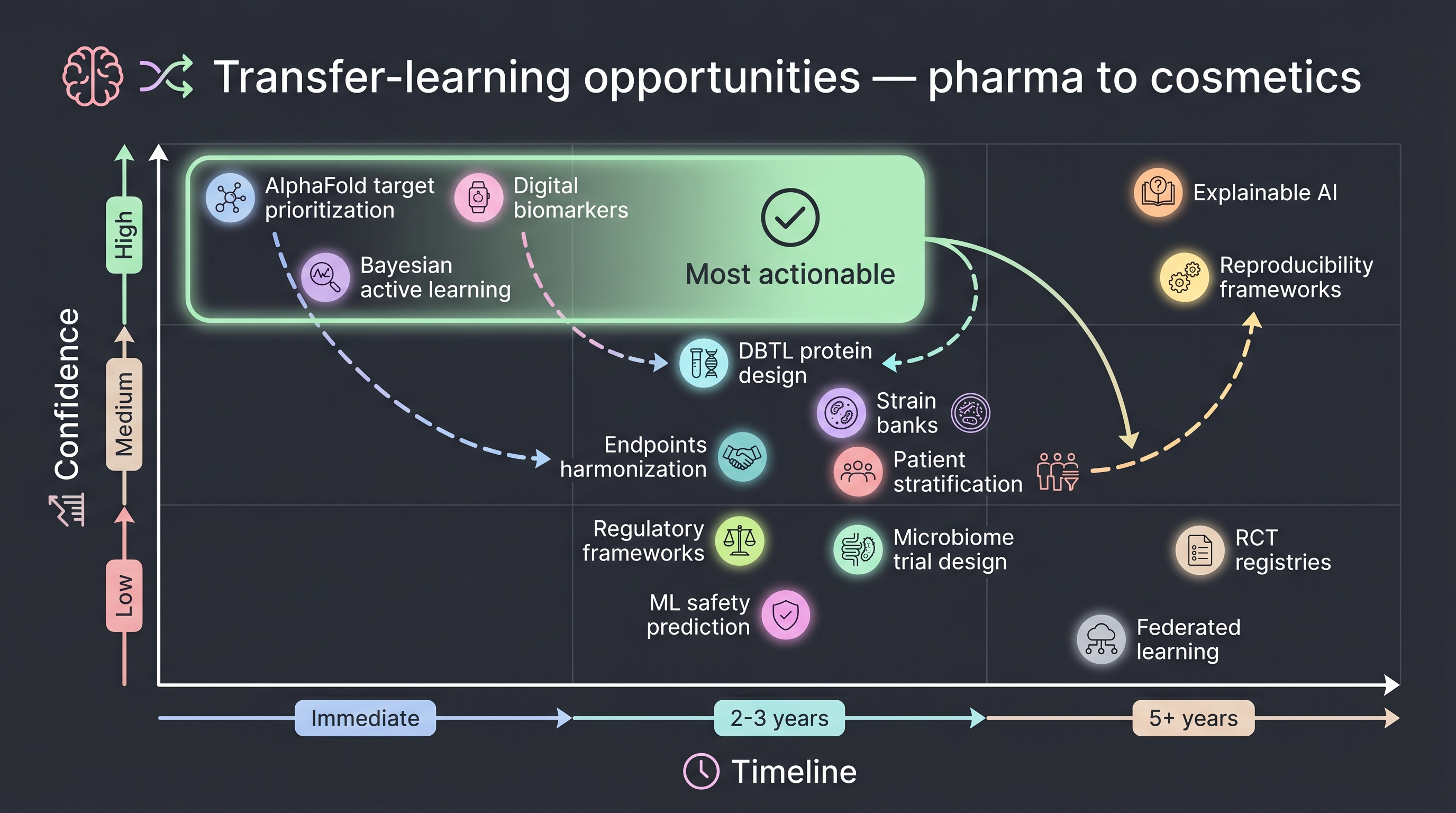
11.9 Transfer-learning matrix — what to borrow from where
The chapter's synthesis — a table a cosmetic R&D planner can use directly in a decision meeting.
| Source industry / case | Cosmetic application | Confidence | Estimated adoption timeline |
|---|---|---|---|
| Insilico PandaOmics + Chemistry42 (target → IND in 30 months) | AI metabolite-target identification + generative active-design pipeline | Medium — pharma scale-up infra missing in cosmetics; renting the SaaS layer is more realistic | 2–3 years (CDMO/OEM-feasible) |
| Insilico Phase 2a publication template (rentosertib +98.4 mL FVC) | The first peer-reviewed clinical-readout publication structure for an AI-designed cosmetic active | High — directly portable. A Korean firm or EU mid-tier could plausibly publish in 2027–2028 | 2–3 years (Gap 1) |
| Seres VOWST ECOSPOR III trial design (binary recurrence endpoint, multi-site) | Clinical-trial design for live-strain or strain-defined postbiotic cosmetic products | Medium — endpoint design and CMC standards are directly portable; capital and regulatory bars differ by orders of magnitude | 3–5 years |
| Vedanta VE303 defined consortium (8 strains, batch consistency) | Cosmetic microbiome-cocktail products — defined N-strain consortia rather than donor-derived | High — Chapter 7's DBTL loop enables it directly; regulatory status remains unresolved | 2–4 years |
| Vedanta VE202 setback (UC indication missed) | Decision frame for not applying the consortium model when the endpoint is not mechanistically microbiome-driven | High — directly portable; pre-validates indication selection | Immediate |
| Cradle Bio SaaS (customer-facing protein design) | Vegan collagen, engineered peptides, metabolite-biosynthesis enzymes designed externally (Chapter 7) | High — Novo Nordisk publicly disclosed as customer; a low-friction path for cosmetic firms | Immediate (PoC within 1 year) |
| Profluent OpenCRISPR-1 (de novo synthetic enzymes) | Microbiome strain engineering — in-house CRISPR tooling or synthetic-enzyme design | Low — weak direct cosmetic use case; SaaS-feasible on a 5-year horizon | 5+ years |
| ESM3 / EvolutionaryScale (foundation protein language model) | Substrate for cosmetic-endpoint fine-tunes (host proteins, microbe-secreted enzymes) | Medium — open-weight release; cosmetic fine-tune know-how stays in-house IP | 2–3 years |
| ZOE PREDICT 1/2 cohort model (100K scale, multi-omics, RCT-backed) | Consumer-facing microbiome-cosmetic personalization with peer-reviewed cohort operation | Medium — capital and cohort-operation scale are infeasible for cosmetic startups; K-beauty's routine-heavy user base offers a partial workaround | 3–5 years |
| Viome metatranscriptomics + B2C model | Marketing pattern of "active function measurement" + consumer-kit logistics | Medium — weak peer-review base implies no reputational moat | Immediate (marketing-only) |
| DayTwo B2B (insurer/provider) model | Cosmetic personalization routed through B2B channels (dermatologists, spas, clinics) | Low — cosmetic retail-direct dominance makes B2B a niche variant | 5+ years |
| MGnify / MIBiG / NPAtlas public databases | Standardizing in-house microbiome/metabolite data, BGC prediction, docking benchmarks | Very high — immediately free; the most underused asset in cosmetic R&D | Immediate |
| Pfizer / Novo / Lilly / GSK AI partnership pattern | In-house R&I + external AI SaaS partner structure; microbiome use case defined internally | High — cosmetic majors (L'Oréal-Microsoft, Shiseido-Accenture) already adopted | Immediate |
| Galderma silence (absence of disclosure) | Market signal — even a global dermatology specialist has not bet publicly; conservative-decision indicator | High — the absence is itself information | Immediate (interpretive) |
How to read the table — the three most reliable items for immediate adoption: (i) public databases (MGnify, MIBiG, NPAtlas), (ii) externalizing protein design via Cradle-style SaaS, and (iii) the Vedanta defined-consortium decision frame. The most reliable item on a 2–3-year timeline: (iv) the Insilico-style Phase 2a publication template — with a plausible scenario of a Korean firm being the first to publish. The clearest non-transferable item: (v) pharma-scale capital intensity and timeline — Insilico's 11-year single-molecule deep dive does not map to cosmetic portfolio economics.
11.10 Open Questions
- Korean cosmetic firms' transfer-learning posture — which items in the table above are already being pursued internally by COSMAX, Amorepacific, and LG H&H, and which remain — publicly — empty? COSMAX's HelloBiome partnership [5] places the personalization layer adjacent to ZOE, but the path to defined consortia or Insilico-style end-to-end publication is not disclosed. Chapter 10 covers part of this — but the question requires direct in-house query.
- First mover in the AI cosmetic clinical-readout race — given Gap 1's vacancy, which firm is most likely to publish first? Korean firms (the COSMAX-Dankook EPI-7 / FACE-LINK continuity [15]), EU mid-tiers (Pierre Fabre, Galderma subsidiaries, the Beiersdorf La Prairie line), or startups (Eligo, Arcaea)? The firm that first applies an Insilico-style end-to-end publication structure to cosmetics is likely to claim defined industrial leadership over the next five years.
- Regulatory convergence between cosmetic and pharma microbiome — when FDA, EU CTR, and Korea MFDS codify a category for live-strain or strain-defined postbiotic cosmetics in the 2027–2030 window, will they pattern after Seres VOWST (BLA) or Geltor PrimaColl (GRAS)? Neither directly applies to cosmetics — a third category is plausible. Chapter 12 treats this as a scenario.
- Nutraceutical skin-gut axis crossover into cosmetics — if ZOE/Viome-type companies begin recording skin endpoints as secondary outcomes in their nutrition data (already framed academically by [30]), combined personalization between oral microbiome supplements and topical cosmetics becomes feasible. A natural pairing with the K-beauty inner-beauty market (collagen drinks, probiotics). But RCT-grade efficacy chains, outside the [10] AHR line, remain weak (Gap 14).
- Cosmetic-specific fork of adjacent data infrastructure — while MGnify, MIBiG, and NPAtlas remain in general microbiome / natural product territory, what is the path to a cosmetic-specific benchmark (cosmetic ingredient × in-vitro skin endpoint, commercial formulation × stability, strain × cosmetic claim)? Industry consortium (KCIA, EFfCI, PCPC), academic consortium (EU H2020), or a single firm partially releasing data (e.g., a slice of the Unilever 30K)? The likeliest first mover is precisely the blueprint question Chapter 12 carries.
References
- Abramson, J., Adler, J., Dunger, J. et al. (2024). Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630:493–500.
- Asnicar, F., Berry, S. E., Valdes, A. M. et al. (2021). Microbiome connections with host metabolism and habitual diet from 1,098 deeply phenotyped individuals. Nature Medicine 27:321–332.
- Asnicar, F., Berry, S. E., Spector, T., Segata, N. (2024). ZOE PREDICT cohort — gut microbiome linked to systemic inflammation and skin-relevant outcomes. PREDICT consortium 2024 papers. [Asnicar et al., 2024]
- Carrieri, A. P., Haiminen, N., Maudsley-Barton, S. et al. (2021). Explainable AI reveals changes in skin microbiome composition linked to phenotypic differences. Scientific Reports 11:4565.
- COSMAX × HelloBiome (2025). Korean Beauty Manufacturer COSMAX × HelloBiome microbiome-powered personalized care. WWD / Personal Care Insights, 2025. [COSMAX × HelloBiome, 2025]
- Cradle Bio (2024). Cradle AI protein design platform — $73M Series B. TechCrunch / SynBioBeta / Tracxn, Nov 26, 2024. [Cradle, 2024]
- Feuerstadt, P., Louie, T. J., Lashner, B. et al. (2022). SER-109, an Oral Microbiome Therapy for Recurrent Clostridioides difficile Infection. New England Journal of Medicine 386:220–229.
- Geltor Inc. (2025). Geltor PrimaColl — first FDA GRAS letter for biodesigned vegan collagen. FoodNavigator-USA / SynBioBeta / Geltor press, Oct 2025. [Geltor, 2025]
- GSK + Imperial College Fleming Initiative (2024). GSK × Fleming Initiative AMR AI programme (£45M). Imperial press 2024; DDW Nov 2025. [GSK × Fleming, 2024]
- Hashimoto-Hachiya, A., Furue, M., Tsuji, G. (2022). Galactomyces Ferment Filtrate Potentiates an Anti-Inflammaging System in Keratinocytes. Journal of Clinical Medicine 11:6338. [Hashimoto et al., 2022]
- Haykal, D., Flament, F., Amar, D. et al. (2025). Cosmetogenomics unveiled: a systematic review of AI, genomics, and the future of personalized skincare. Frontiers in Artificial Intelligence 8:1660356.
- Hayes, T., Rao, R., Akin, H. et al. (2025). Simulating 500 million years of evolution with a language model (ESM3). Science, 15 Jan 2025; preprint bioRxiv 2024.07.01.600583. DOI: 10.1126/science.ads0018.
- Huddleston, J. R. (2014). Horizontal gene transfer in the human gastrointestinal tract: potential spread of antibiotic resistance genes. Infection and Drug Resistance 7:167–176.
- Insilico Medicine clinical authors — Ren, F., Zhavoronkov, A. et al. (2025). A generative AI-discovered TNIK inhibitor for idiopathic pulmonary fibrosis: a randomized phase 2a trial. Nature Medicine, May 2025. [Insilico, 2025]
- Kim, J., Lee, Y. I., Mun, S. et al. (2023). EPI-7-derived ferment as a postbiotic to improve atopic skin. International Journal of Molecular Sciences 24(5):4634. [Kim et al., 2023]
- Lagier, J.-C., Edouard, S., Pagnier, I. et al. (2018). Culturing the human microbiota and culturomics. Nature Reviews Microbiology 16:540–550.
- Louie, T. J., Golan, Y., Khanna, S. et al. (2023). VE303, a defined bacterial consortium, for prevention of recurrent Clostridioides difficile infection: a randomized clinical trial. JAMA 329:1356–1366. [Louie et al., 2023]
- Madaan, T. et al. (2024). Microbiome therapeutics for dermatological conditions — translational landscape. Experimental Dermatology, 2024. [Madaan et al., 2024]
- Mun, S. et al. (2025). FACE-LINK platform — 950-subject Korean facial microbiome × skin phenotype cohort. COSMAX × Dankook University R&D disclosure, 2025. [Mun et al., 2025]
- Nakatsuji, T., Chen, T. H., Narala, S. et al. (2017). Antimicrobials from human skin commensal bacteria protect against Staphylococcus aureus and are deficient in atopic dermatitis. Science Translational Medicine 9:eaah4680. [Nakatsuji et al., 2017]
- Nakatsuji, T., Hata, T. R., Tong, Y. et al. (2021). Development of a human skin commensal microbe for bacteriotherapy of atopic dermatitis. Nature Medicine 27:700–709. [Nakatsuji et al., 2021]
- Novo Nordisk + Cradle (2024). Novo Nordisk × Cradle AI protein design partnership disclosure. Cradle Series B + Novo press, 2024. [Novo Nordisk, 2024]
- Pfizer + XtalPi (2025). Pfizer × XtalPi expanded AI drug discovery collaboration (June 2025). Joint press release, June 2025. [Pfizer × XtalPi, 2025]
- Recursion Pharmaceuticals + Exscientia plc (2024). Recursion and Exscientia merger — formation of integrated AI drug discovery platform. Joint press release, Aug 2024; completion Nov 20 2024. [Recursion × Exscientia, 2024]
- Ren, F., Ding, X., Zheng, M. et al. (2024). A small-molecule TNIK inhibitor (ISM001-055 / rentosertib) discovered via end-to-end generative AI from target identification to Phase 1. Nature Biotechnology, Mar 2024. [Ren et al., 2024]
- Ruffolo, J. A., Nayfach, S., Gallagher, J. et al. (2024). Design of highly functional genome editors by modeling the universe of CRISPR-Cas proteins (Profluent ProGen2 / OpenCRISPR-1). bioRxiv 2024.04.22.590591, Apr 2024. DOI: 10.1101/2024.04.22.590591. [Ruffolo et al., 2024]
- Seres Therapeutics + Nestlé Health Science (2024). VOWST (SER-109) commercial trajectory — first FDA-approved oral microbiome therapeutic. FDA label + Seres/Nestlé HS commercial updates, 2024. [Seres, 2024]
- Unilever Discover Science (2025). Unilever Skin Microbiome and Mental Wellbeing research disclosure. Unilever USA press, 2025. [Unilever, 2025]
- Vedanta Biosciences (2024). Vedanta Biosciences VE303 Phase 3 RESTORATiVE303 trial initiated; pipeline status. Vedanta / PureTech press, May 2024. [Vedanta, 2024]
- Wang, Z., Yuan, F., Zhong, X. et al. (2025). Skin microbiome and skin aging: emerging strategies for manipulation. Microbiological Research 300:128285.
- ZOE PREDICT consortium; Spector, T. et al. (2024). Personalised nutrition by prediction of glycaemic responses to weight loss and microbiome (ZOE METHOD). Nature Medicine, May 2024. DOI: 10.1038/s41591-024-02951-6. [ZOE × Spector, 2024]
- ZOE PREDICT / Asnicar, F. et al. (2025). ZOE Microbiome Health Ranking 2025 — Nature paper on gut microbes associated with health, nutrition and diet. Nature, 2025. DOI: 10.1038/s41586-025-09854-7. [ZOE microbiome ranking, 2025]
- Zhavoronkov, A., Ivanenkov, Y. A., Aliper, A. et al. (2019). Deep learning enables rapid identification of potent DDR1 kinase inhibitors. Nature Biotechnology 37:1038–1040.