Chapter 5: Protein Structure and Interaction Prediction — Efficacy Modeling After AlphaFold
Why this chapter
Chapter 4 covered candidate prioritization — which molecules and strains deserve bench time. This chapter goes one layer deeper: where does the candidate molecule bind on the skin protein, and how, tracked in silico. More than 90% of cosmetic efficacy ultimately reduces to protein–small-molecule interactions: collagen synthesis and degradation (MMP-1, MMP-9), barrier differentiation (filaggrin, AHR), inflammation (NF-κB, AHR, NRF2), pigmentation (tyrosinase, MITF), desquamation (KLK5, KLK7 kallikreins). If you know the geometry and electronics of the binding pocket, you can predict efficacy. If you don't, you regress to brewing a ferment, splashing it onto cells, and waiting — the first-generation method.
Before AlphaFold2's July 2021 release, predicting a single protein structure was a PhD-grade project. The mean GDT-TS of 87.0 (median 92.4) reported by [14] at CASP14 was the first computational result to enter X-ray crystallography's experimental error band (~1 Å). Within four years the cost of predicting one protein structure fell to GPU-minutes, and protein–ligand complex prediction (AlphaFold 3, Boltz-1, Chai-1) became commodity within a single year (2024). This chapter maps the precise topology of that inflection — what it means for cosmetic R&D, and what is still missing.
Three quantitative anchors for this chapter 1. AlphaFold2 reached a median GDT-TS of 92.4 at CASP14 [14] — the first computational model inside crystallographic error. The prior-generation leader sat around GDT-TS 65. 2. Between April and November 2024, AlphaFold 3 → RoseTTAFold All-Atom → Boltz-1 → Chai-1 appeared in sequence, commoditizing protein–ligand complex prediction across closed and open ecosystems [1], [16], [22], [4]. 3. Cosmetic-industry use remains at zero peer-reviewed clinical readouts (Gap 1). The closest adjacent-industry readout is [13]'s rentosertib Phase 2a — 30 months from target to IND. The first cosmetic readout will likely borrow this template (Chapters 9 and 11 follow up).
5.1 Why structure prediction matters in cosmetics
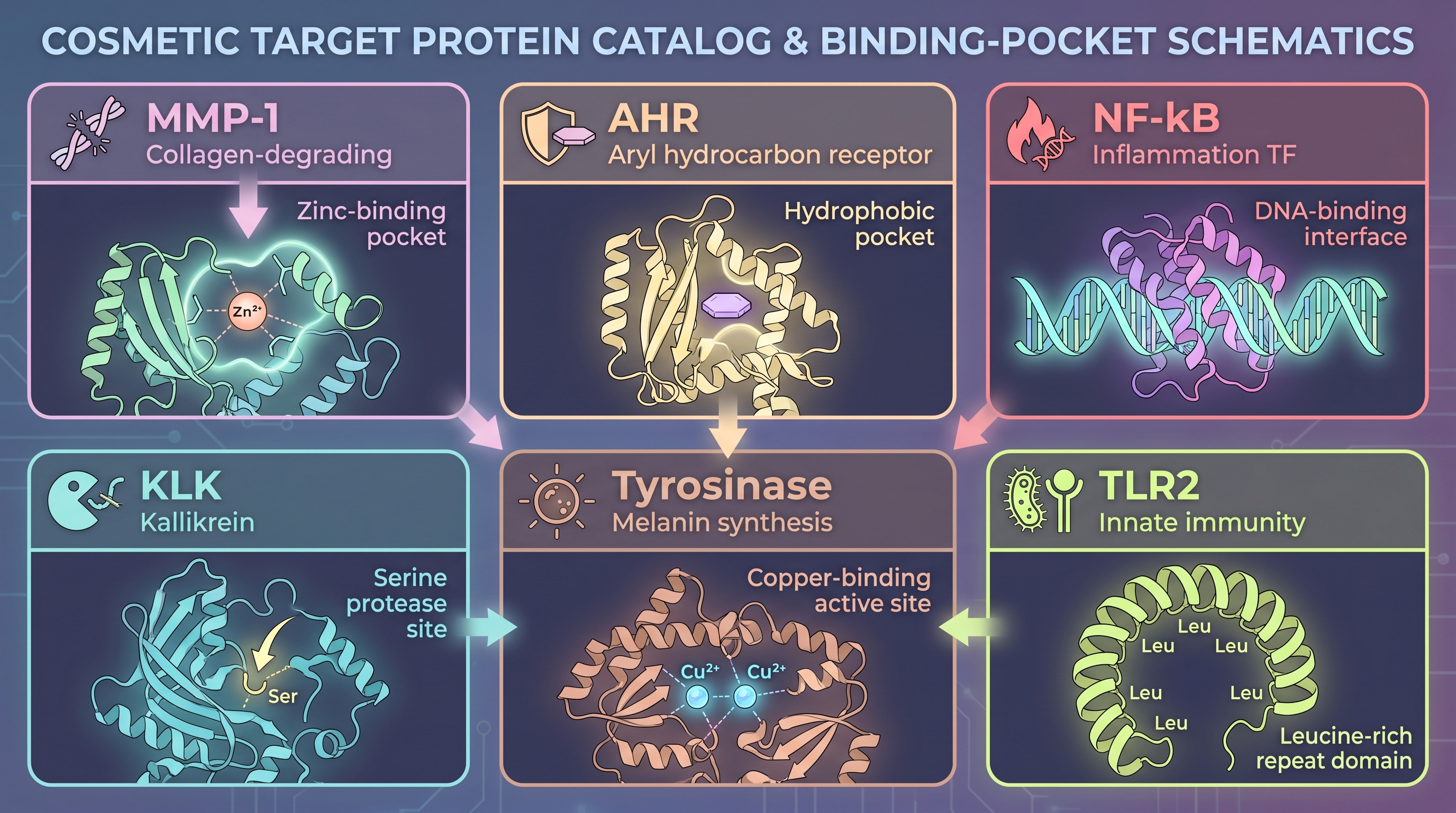
A short tour of the protein roster cosmetic actives target makes the point. Almost every cosmetic R&D catalog converges on the same families.
- Collagen and elastin synthesis axis — COL1A1, COL3A1, ELN, LOX. Synthesis is ribosomal, but blocking degradation is the dominant efficacy mechanism.
- MMP family (matrix metalloproteinases) — MMP-1 (collagenase-1), MMP-3, MMP-9. UV and aging upregulate these enzymes, which cleave collagen. More than half of anti-aging cosmetic actives use MMP-1/9 inhibition as their primary endpoint.
- AHR (aryl hydrocarbon receptor) — co-regulates barrier differentiation (filaggrin, loricrin, involucrin expression) and anti-inflammation. The site where [21] resolved the mechanism of Galactomyces ferment filtrate (SK-II Pitera's source material).
- NF-κB and NRF2 axes — central transcription factors of inflammation and oxidative response. [10] established NRF2 activation as the second molecular pillar of the same Galactomyces filtrate.
- Kallikreins (KLK5, KLK7) — desquamation regulators. Dysregulated in atopic dermatitis and rosacea.
- Tyrosinase / MITF axis — melanin biosynthesis. The core target of brightening endpoints.
- Skin GPCRs and integrins — TLR2, TRPV1, PAR2. Microbial signal recognition and pain/itch sensing.
The structural observation: most of these proteins expose to the cell surface or the extracellular environment. They are reachable by microbial metabolites delivered as cosmetic actives. And if the binding pocket geometry is known, candidate metabolites' binding affinity can be estimated in silico — before the bench. This is one layer deeper than Chapter 4's "molecular fingerprint to endpoint score" pipeline, and it is the new layer AlphaFold made possible.
5.2 AlphaFold2 — the precise location of the inflection
The November 2020 CASP14 (Critical Assessment of protein Structure Prediction) results were the field's pivot. The DeepMind submission described in [14] hit mean GDT-TS 87.0, median 92.4 — a generational gap above second place, and the first computational model inside crystallographic error. The architecture had two innovations:
- Evoformer — jointly updates a multiple sequence alignment (MSA) representation and a pair representation, letting evolutionary and structural information refine each other. Co-evolution between residues hints at spatial proximity, and that proximity in turn refines MSA interpretation.
- Structure module — directly regresses 3D coordinates in an SE(3)-invariant way. Residue frames (N-Cα-C) are expressed as rotations and translations, and Invariant Point Attention (IPA) predicts coordinates, orientations, and confidence (pLDDT) jointly.
Four months later, David Baker's group published RoseTTAFold [3] — a three-track (sequence, distance, coordinates) neural network that matched AlphaFold2 accuracy at one-fifth the compute. The simultaneous arrival was not coincidence: Evoformer-class architectures were the field's convergence point, not one group's miracle.
Industry effect was immediate. [17]'s ESMFold (Meta AI / EvolutionaryScale lineage) released 617 million predicted metagenomic protein structures in the ESM Metagenomic Atlas during 2022–2023. Because ESMFold infers structure from language-model embeddings alone — no MSA — it runs 60× faster than AlphaFold2. For cosmetic R&D, this means "almost every public microbial metagenomic protein is already structurally annotated." iHSMGC's 10.94 million skin-microbiome genes (Chapter 3) can be assumed to have coordinates after one ESMFold pass.
AlphaFold2's limits were equally clear. (1) It was tuned for single-chain monomers — complexes only stabilized after [6]'s AlphaFold-Multimer. (2) It could not handle ligands, ions, DNA, or RNA — drug-metabolite docking required separate tools (AutoDock Vina, GNINA, etc.). (3) Dynamic conformations, allosteric states, and lipid-protein interactions in membrane proteins fell outside its training scope.
The third limitation stings for cosmetics. Many cosmetic targets are not secreted enzymes like MMPs but membrane-spanning receptors (TRPV1, PAR2, integrins). AlphaFold2's single-structure output cannot cleanly separate ligand-bound active state from ligand-free inactive state, nor does it represent lipid-bilayer context. AlphaFold 3 only partially closed this gap.
5.3 AlphaFold 3 — metabolite-protein complexes without crystals
[1]'s AlphaFold 3, published in Nature in May 2024, extended structure prediction from "single protein" to whole biomolecular systems. Proteins, DNA, RNA, ligands, ions, and a subset of covalent modifications (phosphorylation, some glycosylation) live in one model. Two architecture changes drive this:
- Diffusion-based coordinate generation — the original structure module is replaced by a diffusion module that refines coordinates from noise. This is the same paradigm as RFdiffusion and de novo protein design pipelines, and the advantage is uniform handling of diverse molecular types.
- Comprehensive input encoding — proteins, small-molecule SMILES, nucleic-acid sequences, and ions enter the same grammar. From the user's perspective, "complex of protein X + ligand Y" is a single request.
On PoseBusters, AlphaFold 3 cleared 76.4% of protein–ligand poses, well above AutoDock Vina's ~52%. It also outperformed RoseTTAFoldNA on protein–DNA and protein–RNA complexes. Cosmetic implication: the binding pose of, say, MMP-1 with a microbial metabolite (spermidine from [15]'s COSMAX work, short-chain fatty acids from Cutibacterium, porphyrins) can be predicted without a crystal. The IC50 of a collagen-degradation inhibitor can be estimated before the bench.
[1] also names two boundaries. (1) Affinity (ΔG, Kd, IC50) is not directly predicted — only the pose. Quantitative affinity requires post-processing (MM-PBSA, FEP, or an ML scoring function). (2) Commercial license was a sticking point: at release the model was accessible only via the DeepMind AlphaFold Server for non-commercial use. Weights for non-commercial academic use were released between November 2024 and early 2025 (the [1] license supplement), leaving industrial use in a gray zone. That license friction is what lit the fuse on open replication.
5.4 The open-source replication race — Boltz, Chai, RoseTTAFold-AllAtom, ESM3
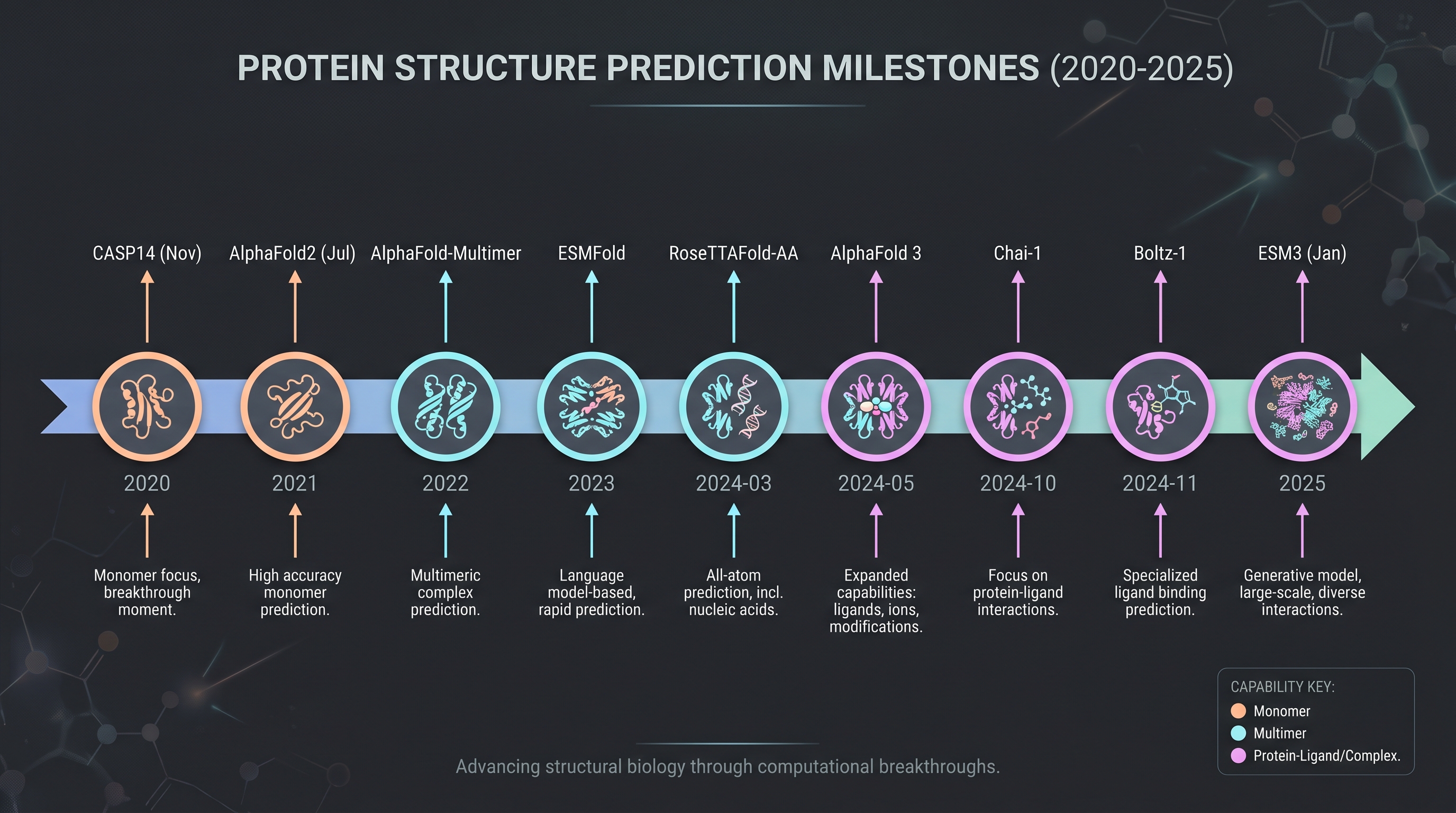
AlphaFold 3's closed posture motivated academia and adjacent industry. Within 2024, three open-weight (or near-open) replication models appeared.
RoseTTAFold All-Atom [16] (Science, March 2024). The Baker group's successor — three-track networks generalized to all-atom representation. Proteins plus nucleic acids, small molecules, and metal ions in one model. Its natural composability with RFdiffusion makes de novo protein design part of the same pipeline. Weights are open for non-commercial academic use. Cosmetic relevance: scenarios like designing collagen-mimic peptides that competitively inhibit MMPs become feasible — connecting directly to Chapter 7's synthetic biology.
Boltz-1 [22] (MIT Jameel Clinic technical report, November 2024). The closest open-weight replication of AlphaFold 3. MIT Jameel Clinic's OpenAlphaFold-3 initiative released it on November 19. The MIT license permits commercial use — decisive for cosmetic and bio industries that wanted to download weights and fine-tune in-house. PoseBusters accuracy lags AlphaFold 3 by 1–5%, but licensing freedom offsets the gap. The lesson: industrial adoption thresholds are licensing, not raw accuracy.
Chai-1 [4] (October 2024). The Chai Discovery startup's AlphaFold 3-equivalent model. Non-commercial academic weights plus a commercial API. Self-reported benchmarks claim parity or slight wins over AlphaFold 3 on protein–ligand pose subsets. The SaaS licensing model presents a different trade-off from Boltz — no operational overhead, but data and results flow outside the firm.
ESM3 [12] (Science, January 2025). EvolutionaryScale's protein language model. 98B parameters total (a 7B-parameter version is open), trained to unify protein sequence, structure, and function as a single token stream. It is the evolution of ESMFold (2022–2023) but generates function and sequence in addition to structure — that is, "design a new protein sequence that fits this binding pocket" is a supported prompt. EvolutionaryScale launched in June 2024 with $142M seed funding plus AWS and NVIDIA infrastructure partnerships [7]. ESM3 is the most directly applicable tool from a cosmetic and synbio perspective. Chapter 7's vegan collagen (Modern Meadow / Evonik Vecollage [8], Geltor PrimaColl [9]) is the first industrial readout of proteins designed with this class of tool.
The simultaneous arrival of these four models tells a simple story: AlphaFold 3-class tools are commodity as of late 2024. The choice is licensing (closed SaaS, open academic, open commercial), not accuracy — the spread is 1–5%. The implication for cosmetic R&D: weight access is no longer the barrier to entry.
[2]'s tool, published in Bioinformatics Advances, responds to this convergence — it runs AlphaFold 3, Boltz-1, and Chai-1 on the same input and quantitatively compares results. Industry standard is consolidating around "run all three and inspect consensus."
5.5 A cosmetic-target docking pipeline — concrete walk-through
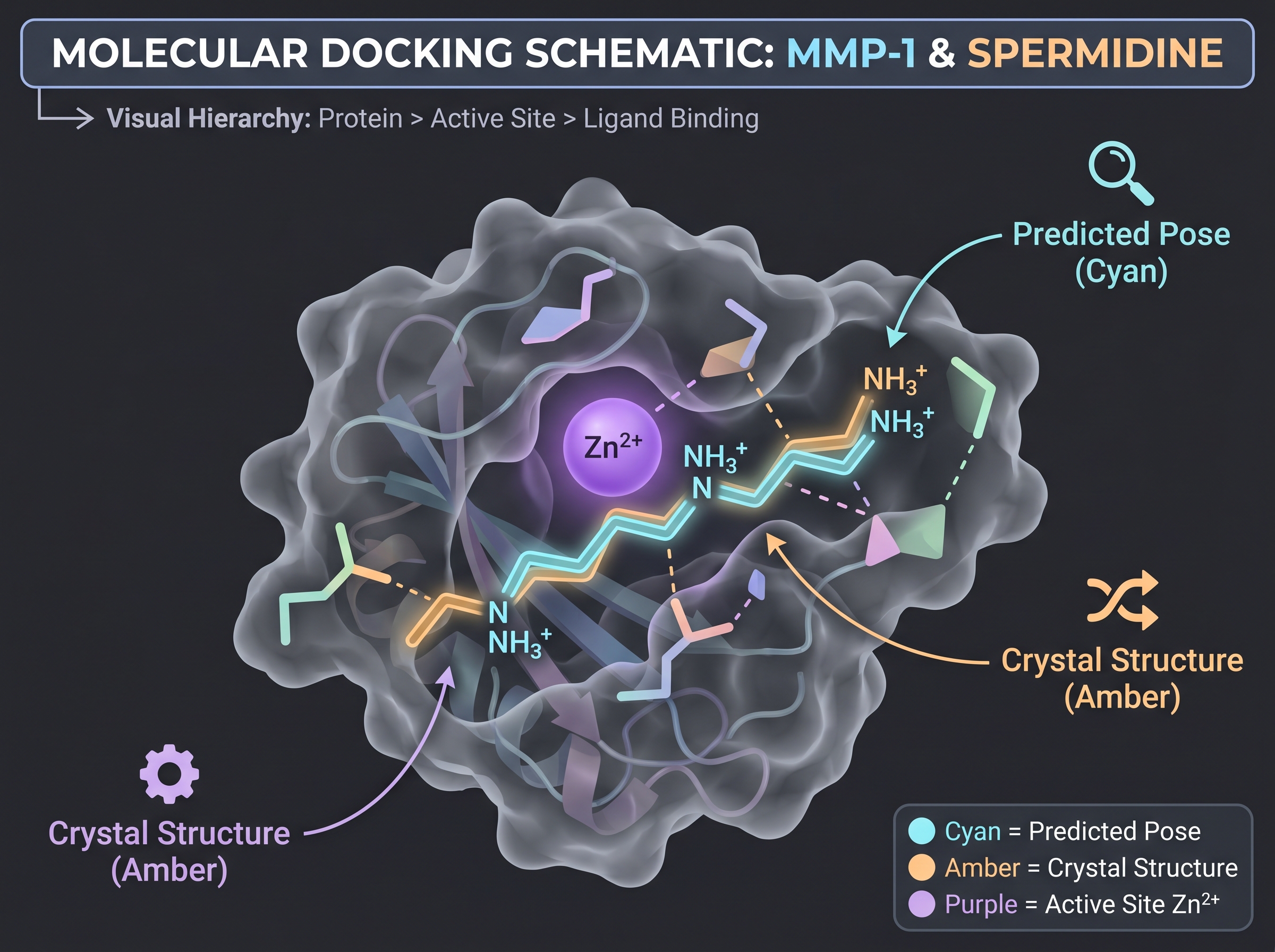
To anchor the abstraction, follow one microbial metabolite end-to-end through the pipeline. The example is spermidine — a polyamine produced by skin microbes including Lactobacillus — published by the COSMAX/Dankook group in Communications Biology [15].
Step 1 — collect structural inputs.
- Spermidine is a small molecule, encoded as SMILES
NCCCCNCCCN. Available in ChEMBL and PubChem. - Candidate skin protein targets: MMP-1, MMP-9 (collagen degradation), TGM3 (transglutaminase, cornified envelope cross-linking), AHR. [15] reported fibroblast collagen recovery in vitro, so collagen synthesis-degradation is the most plausible axis.
- MMP-1 structure: PDB ID 1HFC, 1.56 Å X-ray. Also indexed in AlphaFold DB under UniProt P03956.
Step 2 — docking.
- Submit "MMP-1 (UniProt P03956) + spermidine (SMILES)" to AlphaFold 3 or Boltz-1. Output: binding pose plus per-residue confidence (pLDDT). Core question: does spermidine sit near the zinc-coordinating catalytic site of MMP-1, or in an allosteric pocket?
- Baseline: submit the same input to AutoDock Vina. AlphaFold 3-class poses typically achieve RMSD within 1–2 Å of the crystal pose ([1] Figure 4).
Step 3 — affinity estimation.
- AlphaFold 3 and Boltz-1 produce poses, not ΔG. Post-process with MM-GBSA or MM-PBSA to estimate ΔG — typically ±2 kcal/mol, which fixes IC50 to the right order of magnitude.
- Alternatively, an ML scoring function (GNINA, DiffDock-L).
- Example output: estimated IC50 ~50–200 μM — a range that needs bench confirmation.
Step 4 — wet-lab validation (the "AI predicts, culture validates" thesis from Chapter 3).
- MMP-1 enzymatic activity assay (EnzChek fluorogenic substrate): measure spermidine IC50.
- Primary fibroblast: procollagen ELISA, MMP-1 Western blot.
- Compare predicted vs measured IC50. If the predicted and measured values land within the same order of magnitude, the candidate passes. If they differ by more than 10×, reinspect the binding-pocket assignment.
Step 5 — accumulate, generalize.
- Validated poses feed back into internal training data. Subsequent candidates (Cutibacterium short-chain fatty acids, Staphylococcus epidermidis epifadin peptides) enter the docking pipeline with this prior baked in.
This five-step pipeline is identical in shape to [13]'s journey from TNIK target to IPF Phase 2a — the only differences are cosmetic vs drug endpoint and industry incentive.
5.6 Limits and remaining gaps
Even with commodity tools, several cosmetic-domain bottlenecks remain. An honest map:
5.6.1 Affinity quantification is much harder than pose prediction.
[1] explicitly notes that AlphaFold 3-class models predict pose well but do not predict affinity directly. The post-docking ΔG estimators (MM-PBSA, FEP, ML scoring functions) all have failure modes, and ±2 kcal/mol error is typical — about one order of magnitude on IC50. From a cosmetic R&D perspective, "IC50 50 μM" versus "IC50 500 μM" decides whether the candidate enters formulation — pose alone cannot draw that line.
5.6.2 PTMs are common on cosmetic targets.
Many skin proteins carry glycosylation (N-glycosylation), phosphorylation, or sulfation. MMP-9 activity is phosphorylation-dependent, and epidermal proteins (filaggrin, involucrin) carry extensive glycosylation. AlphaFold 3 explicitly handles some PTMs but not all, and complex glycan chains relevant to cosmetic targets are still imperfectly modeled ([1] supplementary).
5.6.3 Membrane-spanning receptors — GPCRs and integrins.
Most cosmetic sensing targets (TLR2, TRPV1, PAR2, integrin αVβ3) are membrane-spanning. AlphaFold-class models do not explicitly model lipid bilayer environment, and active versus inactive GPCR conformations are not cleanly distinguished. RoseTTAFold-AllAtom [16] and ESM3 [12] partially improve this, but ligand-pose accuracy on membrane targets still trails cytosolic proteins. Cosmetic R&D on GPCR targets therefore needs more conservative interpretation of docking outputs.
5.6.4 Dynamic conformations and allosteric sites.
A single static structure cannot capture conformational ensembles. MMP-3 toggles between open and closed states with different ligand affinities. Workarounds — running molecular dynamics (GROMACS, OpenMM) as post-processing, or running AlphaFold-Multimer with alternative MSAs to force ensemble generation — exist but are not standardized.
5.6.5 The validation-data void (Gap 1).
[11]'s PRISMA review of 74 cosmetogenomics-AI studies explicitly reports zero peer-reviewed clinical readouts of AI-designed cosmetic actives. Even with commodity tools, if industry does not publish clinically validated products built with them, the models' external validity remains untested. That is the direct cause of cosmetic-industry credibility lagging pharma in academic comparison.
5.7 The opportunity gap — Insilico's readout needs a cosmetic twin
[11] is explicit: as of May 2026, no peer-reviewed clinical readout exists for an AI-designed cosmetic active. The closest adjacent readout is [13]'s Nature Medicine rentosertib trial — TNIK inhibitor for idiopathic pulmonary fibrosis (IPF), Phase 2a, 12-week +98.4 mL FVC versus placebo −20.3 mL. [19]'s companion Nature Biotechnology paper documents 30 months from target identification (PandaOmics) to IND — versus the traditional 6–8 years.
Templates a cosmetic firm could borrow:
- AI for target identification — causal proteins for skin aging (multi-omics target ID via tools like PandaOmics, paired with AlphaFold structural availability filters).
- Molecular design — generative chemistry (Chemistry42, Cradle [5], Profluent [20]) plus AlphaFold 3-class docking validation.
- Compressed synthesis and validation — synbio strains (Chapter 7) plus ex vivo skin models (Chapter 9).
- Clinical readout publication — Insilico's pattern of dual publication in Nature Biotechnology and Nature Medicine has become a credibility standard. A cosmetic firm publishing in equivalent channels (JAAD, British Journal of Dermatology, Skin Pharmacology and Physiology) would create a category-defining asset.
Most plausible first-readout candidates: a Korean firm (COSMAX, Amorepacific, LG H&H) or a smaller EU player (Lactobio-style microbiome specialists). The Western Big-5 (L'Oréal, Unilever, Estée Lauder, P&G, Shiseido) more likely default to press releases — marketing ROI usually beats peer-reviewed publication. Industry mapping in Chapters 10 and 11.
Vegan collagen is the first step taken. [9]'s PrimaColl received the first FDA GRAS "no questions" letter for biodesigned vegan collagen in October 2025, and [8]'s Vecollage Fortify L (in partnership with Modern Meadow) became the first cosmetic-grade vegan collagen launch. Both reportedly used ESM3- and RoseTTAFold-AllAtom-class tools for protein design — but neither has a peer-reviewed clinical readout; the visible evidence stops at GRAS letters and press releases.
5.8 Open Questions
- A step change in affinity prediction — what data-architecture combination compresses ±2 kcal/mol error to ±0.5 kcal/mol? FEP plus ML scoring fusion is the most plausible near-term answer, but cosmetic-domain affinity data is far sparser than pharmaceutical equivalents in ChEMBL.
- Closing the membrane-protein gap — AlphaFold 3-class tools consistently underperform on GPCR active/inactive separation and lipid-protein modeling. Because many cosmetic sensing targets are GPCRs, this gap has direct business impact. Which tools and datasets will close it?
- Cost asymmetry of the simulation–validation cycle — docking has dropped to GPU-minutes per candidate, but bench IC50 confirmation remains $10s–$100s per candidate. Where does this asymmetry move the bottleneck in cosmetic R&D? Chapter 9 extends this question to the clinical stage.
- In silico handling of PTMs — complex glycosylation and sulfation common to cosmetic targets remain partially modeled in AlphaFold 3. What tooling will close the gap? RoseTTAFold-AllAtom plus GLYCAM-class sugar-binding tools is a candidate path.
- Implications of the data asymmetry — as long as cosmetic R&D withholds docking results from publication, model external validity depends on pharmaceutical readouts. How does that dependence shape cosmetic tool selection and validation standards? Chapter 12 integrates the answer.
References
- Abramson, J., Adler, J., Dunger, J. et al. (2024). Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500.
- ABCFold authors (2025). ABCFold: easier running and comparison of AlphaFold 3, Boltz-1, and Chai-1. Bioinformatics Advances 5(1):vbaf153. [ABCFold, 2025]
- Baek, M., DiMaio, F., Anishchenko, I. et al. (2021). Accurate prediction of protein structures and interactions using a three-track neural network. Science 373(6557):871–876.
- Chai Discovery Team (2024). Chai-1: Decoding the Molecular Interactions of Life. bioRxiv 2024.10.10.615955; Chai Discovery technical report. [Chai Discovery, 2024]
- Cradle Bio (2024). Cradle AI protein design platform — $73M Series B. TechCrunch / SynBioBeta, Nov 2024. [Cradle, 2024]
- Evans, R., O'Neill, M., Pritzel, A. et al. (2022). Protein complex prediction with AlphaFold-Multimer. bioRxiv 2021.10.04.463034.
- EvolutionaryScale (2024). EvolutionaryScale launch with ESM3 — $142M seed and commercial protein-design platform. Press release, June 25, 2024. [EvolutionaryScale, 2024]
- Evonik (2024). Evonik Vecollage Fortify L — vegan collagen for cosmetic personal care (Modern Meadow partnership). Evonik press, Feb 2024. [Evonik, 2024]
- Geltor (2025). Geltor PrimaColl — first FDA GRAS letter for biodesigned vegan collagen. FoodNavigator-USA / SynBioBeta / Geltor press, Oct 2025. [Geltor, 2025]
- Hashimoto, K., Yamamoto, T., Yagi, M. et al. (2022). NRF2 activation by Galactomyces ferment filtrate complements the AHR-axis mechanism in skin barrier homeostasis. Journal of Cosmetic Dermatology, 2022. [Hashimoto et al., 2022]
- Haykal, D., Flament, F., Amar, D. et al. (2025). Cosmetogenomics unveiled: a systematic review of AI, genomics, and the future of personalized skincare. Frontiers in Artificial Intelligence 8:1660356.
- Hayes, T., Rao, R., Akin, H. et al. (2025). Simulating 500 million years of evolution with a language model (ESM3). Science, Jan 2025.
- Insilico Medicine clinical authors, Ren, F., Zhavoronkov, A. et al. (2025). A generative AI-discovered TNIK inhibitor for idiopathic pulmonary fibrosis: a randomized phase 2a trial. Nature Medicine, May 2025. [Insilico, 2025]
- Jumper, J., Evans, R., Pritzel, A. et al. (2021). Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589.
- Kim, G., Kim, M., Kim, M. et al. (2021). Spermidine-induced recovery of human dermal structure and barrier function by skin microbiome. Communications Biology 4:231.
- Krishna, R., Wang, J., Ahern, W. et al. (2024). Generalized biomolecular modeling and design with RoseTTAFold All-Atom. Science 384:eadl2528.
- Lin, Z., Akin, H., Rao, R. et al. (2023). Evolutionary-scale prediction of atomic-level protein structure with a language model (ESMFold + ESM Metagenomic Atlas). Science 379:1123–1130.
- Novo Nordisk; Cradle (2024). Novo Nordisk × Cradle AI protein design partnership disclosure. Cradle Series B + Novo press, 2024. [Novo Nordisk, 2024]
- Ren, F., Ding, X., Zheng, M. et al. (2024). A small-molecule TNIK inhibitor (ISM001-055 / rentosertib) discovered via end-to-end generative AI from target identification to Phase 1. Nature Biotechnology, Mar 2024.
- Ruffolo, J. A., Nayfach, S., Gallagher, J. et al. (2024). Design of highly functional genome editors by modeling the universe of CRISPR-Cas proteins (Profluent ProGen2 / OpenCRISPR-1). bioRxiv 2024.04.22.590591.
- Takei, K., Mitoma, C., Hashimoto-Hachiya, A. et al. (2015). Galactomyces ferment filtrate as AHR agonist restoring filaggrin and skin barrier function. Journal of Dermatological Science, 2015. [Takei et al., 2015]
- Wohlwend, J., Corso, G., Passaro, S. et al. (2024). Boltz-1: Democratizing Biomolecular Interaction Modeling. bioRxiv 2024.11.19.624167; MIT Jameel Clinic technical report.