Chapter 3: NGS and Metagenomics — How We Read the Microbiome
Why this chapter. The single biggest reason cosmetic microbiome moved from the fermentation era to the data era is that the cost of reading a microbe fell roughly five orders of magnitude over thirty years. Over the same window, the craft of growing a microbe did not disappear — it returned under the banner of culturomics and is now the natural complement to sequencing. This chapter walks the reading technologies (16S, ITS, shotgun, long-read) and the culturomics revival as one continuous story, then states the book's position on Brief A's central question: are fundamental wet-lab methods — isolation, identification, culture, antimicrobial assays — now obsolete? No. They are complementary. AI predicts; culture validates.
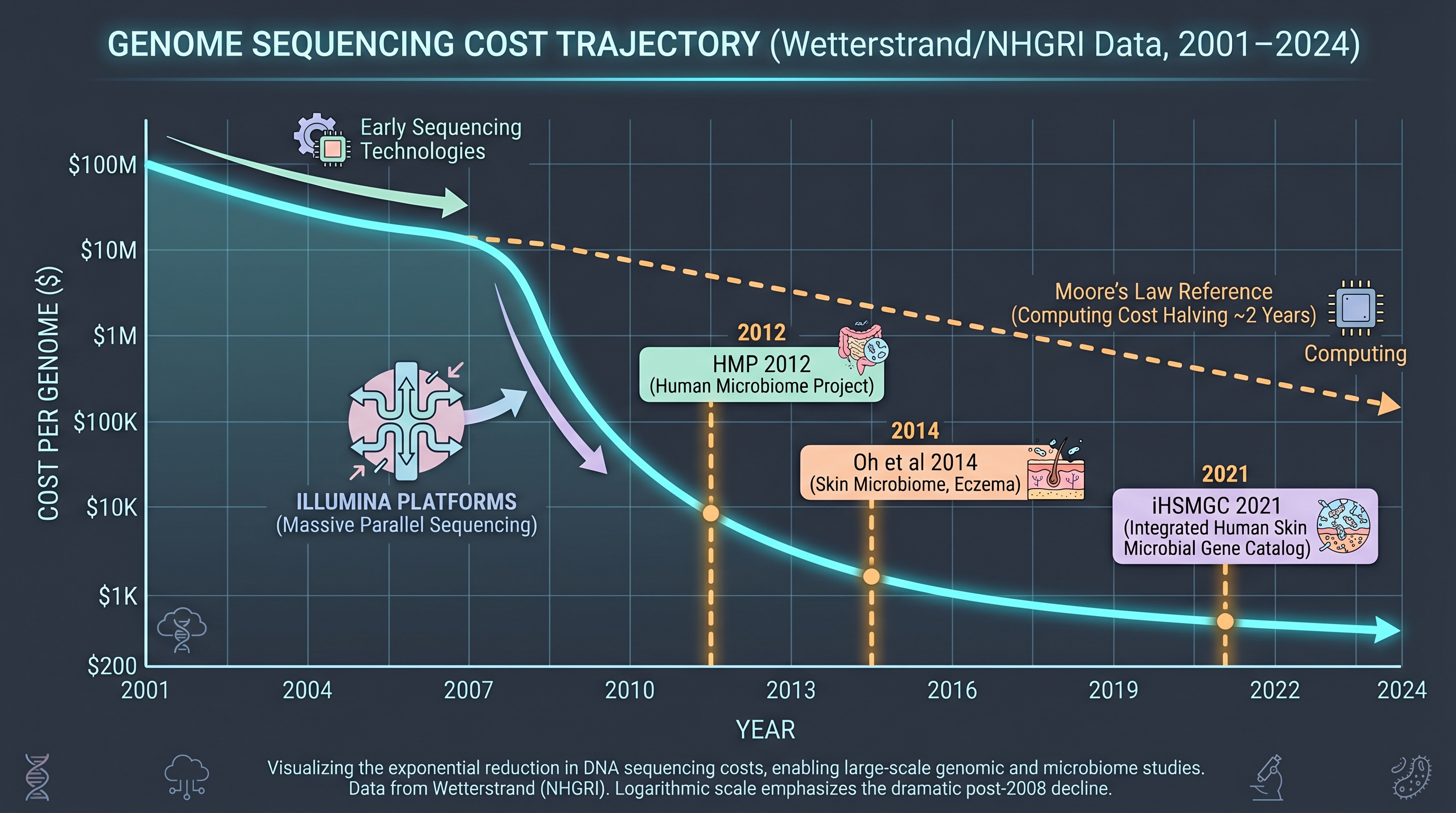
Three quantitative anchors. - The cost of sequencing one human genome fell from ~$100M in 2001 to ~$200 in 2024 — roughly 5×10⁵ ×. This curve is the substrate of every downstream industrialization story in the book. - Industrial consortia (L'Oréal, Unilever, COSMAX) hold cohorts of tens to hundreds of thousands of samples, but the public HMP reference covered only 3 skin sites and the iHSMGC catalog found ~45% of skin genes that HMP missed in an East-Asian cohort. Data asymmetry is itself a chapter-12 starting line. - Culturomics had recovered >1,000 previously uncultured human-associated species by 2018 — yet a skin-specific equivalent catalog at gut scale does not exist. This is one of the book's named gaps.
3.1 Why We Needed to Read — From Microscope to Genome
The first eyes on a skin microbe were Antonie van Leeuwenhoek's homemade microscope, late 17th century. Two more centuries — Robert Koch's plates, Pasteur's broths — taught us to isolate, grow, phenotype. Fermented cosmetics are an heirloom of that era (Chapter 1). But by the late 1990s two things had become clear.
First, most skin microbes do not grow on standard media — the great plate count anomaly. Second, within a single species, strain identity routinely determines whether a microbe is friend, neighbor, or pathogen. The 16S rRNA gene as a taxonomic clock (and the sequencing to read it) opened a road around culture: see them without growing them.
Two infrastructures made the road real. One was the next-generation sequencing (NGS) cost curve — from $100M per genome in 2001 to ~$200 by 2024. The other was the standardization of open-source analysis pipelines. [5] released QIIME, which has accrued over 40,000 citations and made it possible for any lab to process amplicon data the same way; [2] QIIME 2 is its provenance-tracked, plugin-based successor and is now the academic default.
3.2 16S rRNA Amplicon — A Cheap First Telescope
The 16S rRNA gene is universal in bacteria (conserved regions) and species-discriminating (hypervariable regions). The dominant practice is to PCR-amplify a variable region — most often V3–V4 — and read it on short-read Illumina. Costs are low enough to scan thousands of samples per cohort.
[5] standardized OTU (Operational Taxonomic Unit, 97% similarity) clustering. But 97% OTUs sit closer to genus-leaning species than to true species — biologically distinct strains routinely fell into the same OTU. The break came with [4] DADA2, which models Illumina's error process and resolves Amplicon Sequence Variants (ASVs) at single-nucleotide resolution. Strain-level diversity that had been collapsed under 97% OTUs became visible for the first time.
Three reference databases — [17] SILVA, Greengenes, and RDP — split the taxonomy backbone. Every taxonomic renaming (e.g., Propionibacterium acnes → Cutibacterium acnes) propagates at different speeds across these references, and that desynchronization is a small but persistent source of cross-study irreproducibility.
Limits. 16S amplicon has three structural weaknesses. (i) Region choice (V1–V9) matters: Staphylococcus resolves better in V1–V3; Lactobacillus in V3–V4. (ii) Fungi and viruses are invisible — ITS and viromics are required separately. (iii) Most fundamentally, 16S tells you who is there, not what they do. For an efficacy hypothesis, that is not enough.
3.3 ITS and the Fungi — A Separate Telescope for Malassezia
Skin fungi do not show up in 16S. [6] performed the first systematic ITS1 survey of the skin mycobiome alongside bacterial 16S across 14 sites in 10 healthy adults. The result was striking: Malassezia dominates 11 core-body and arm sites (often >80% of fungal reads), whereas the foot harbors a much more diverse mycobiome including Aspergillus, Cryptococcus, and Rhodotorula.
Malassezia matters cosmetically: dandruff, seborrheic dermatitis, and a portion of atopic-dermatitis endpoints track Malassezia species composition (Chapter 2). ITS profiling is therefore effectively mandatory for antifungal / anti-dandruff efficacy work. ITS1 is a single locus and under-resolves some Malassezia species; frontier studies pair it with ITS2 or use long-read full-rRNA reads as the structural fix.
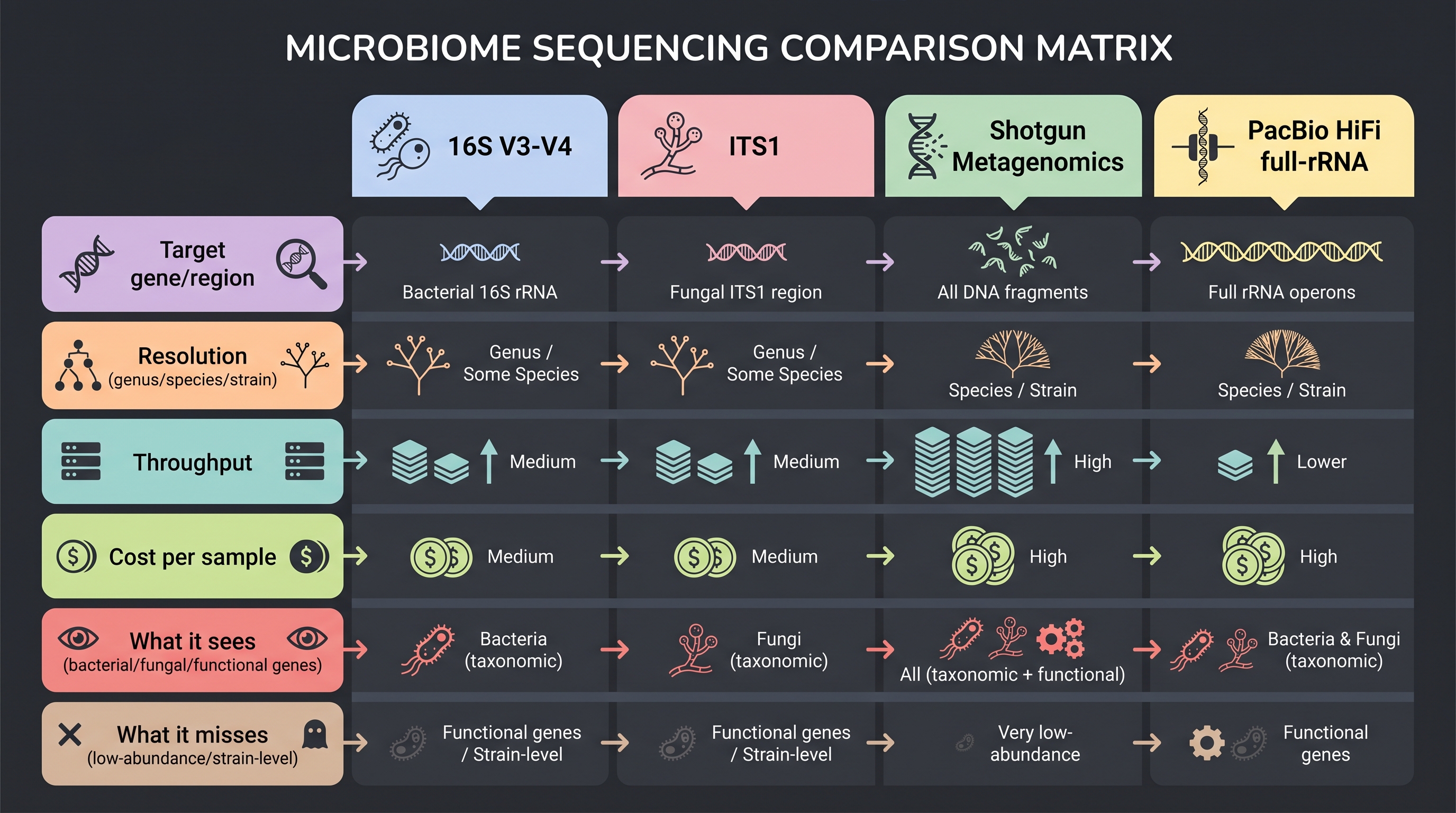
3.4 Shotgun Metagenomics — A Second Telescope That Sees Function
Shotgun metagenomics randomly fragments and sequences all DNA in a community. If amplicon is a taxonomic photograph, shotgun is closer to a functional blueprint. The catch is real: skin is low-biomass and >90% host DNA, so adequate microbial coverage demands ~10 Gb per sample — 10–50× the cost of 16S.
Skin shotgun became practical with [14]. The NIH NHGRI Segre lab assembled 270 metagenomes (15 adults × 18 sites) and produced the first high-coverage strain-level reconstruction across bacteria, fungi, and viruses on healthy skin. The follow-up [15] resampled the same individuals 1–2 years later and — surprisingly — found median strain stability >80% at sebaceous sites over 2 years. Skin microbiomes are far more stable than the "environmental exposure" intuition suggests. This single finding is the scientific bedrock of "restore your microbiome balance" marketing and also the warning that products struggle to durably shift composition.
The analysis tooling kept pace. [18] MetaPhlAn2 uses clade-specific marker genes for fast species-level profiling, and its companion StrainPhlAn module extends to strain-level tracking. [1] HUMAnN reconstructs KEGG/MetaCyc pathway abundance from reads — the first quantitative input to any predict-metabolite-production-from-sequencing workflow used in cosmetic screening.
iHSMGC and data asymmetry. Where the reference catalog gets built shapes what gets seen. [11] iHSMGC combined 822 Han Chinese shotgun samples with 538 prior North American samples and assembled a 10.9 M-gene skin catalog, ~4.88 M (≈45%) of which were absent from Western references. The same study defined two "cutotypes" on East-Asian facial skin: a Moraxella osloensis-type and a Cutibacterium acnes-type — and M. osloensis is nearly absent in the North-American cohort.
iHSMGC carries a two-layer message. On the surface: "Western reference cohorts under-count nearly half the gene content of East-Asian skin." Underneath: current public references have systematic blind spots determined by where and on whom they were built — and AI efficacy models trained on these references inherit that bias by construction. We pick this thread back up in (Chapter 4) on AI screening and (Chapter 12) on infrastructure.
3.5 Long-Read — Strain Resolution and Hybrid Assembly
Illumina short-reads (150–300 bp) hit walls at repetitive regions and at distinguishing closely related strains. Long-read platforms — PacBio HiFi (>15 kb, Q30+) and Oxford Nanopore (kb to Mb) — solve both problems by carrying more information per read. Full 16S (~1,500 bp) and the 16S–23S ITS region fit inside a single read, giving species- to strain-level resolution even on amplicons; on shotgun, plasmids, repeats, and mobile elements assemble intact.
Cost-sensitive cohorts still default to Illumina + DADA2, but cosmetic R&D that needs strain-level precision — for example, (Chapter 7) synbio strain engineering or (Chapter 4) AI strain screening's ground-truth — has effectively standardized on long-read + short-read hybrid assembly. Costs fell enough by 2024 that a handful of Korean and Chinese industrial cohorts now run it routinely.
3.6 The Culturomics Revival — "Culture Is Dead" Was Premature
A brief consensus right after NGS arrived held that culture was over. The most decisive rebuttal is [10], the Nature Reviews Microbiology culturomics synthesis. The Marseille IHU group combined 70+ media, varied gas mixtures, and MALDI-TOF MS identification to recover over 1,000 previously uncultured human-associated species by 2018, many of which had been known only as 16S signals before.
Why did culture come back? Three reinforcing reasons.
First, functional validation. 16S and shotgun tell you that something is or could be there. Measuring antimicrobial, anti-inflammatory, or anti-aging activity on a cosmetic-relevant endpoint requires the organism in hand. AI may recommend a metabolite candidate, but only a held strain that you can grow, ferment, and extract can confirm efficacy in vitro (Chapter 4, Chapter 5).
Second, strain banks as assets. Synthetic biology era R&D treats strains as IP (Chapter 7). Sequences do not own strains — culture collections do. Beyond public banks (ATCC, DSMZ, KCTC), proprietary internal collections at COSMAX, Amorepacific, and Galderma are core competitive moats.
Third, ground truth for AI. An AI model that predicts "this strain is anti-inflammatory" can be trusted only if (a) ground-truth labels covered a sufficiently diverse input distribution and (b) new predictions can be validated quickly. High-throughput culturomics is the infrastructure that delivers both. AI predicts; culture validates. That one line is the book's load-bearing thesis.
But skin culturomics is still thin. Most of Lagier's 1,000+ recoveries are gut-derived. Skin-specific culturomics lags because of (a) host-DNA identification noise, (b) low biomass, and (c) the demanding media — oxygen, CO₂, lipid dependence — that skin commensals often need. The frontier corpus contains essentially no skin-specific culturomics paper beyond [10]. This is the book's Gap 3, and it weakens the foundation under (Chapter 9) clinical validation.
The hardest counterexample is the Gallo-lab lineage at UCSD. [12] showed that healthy-skin commensal staphylococci — particularly S. hominis and S. epidermidis — secrete antimicrobial peptides that kill S. aureus, and that this capacity is deficient in atopic-dermatitis patients. [13] then took a single isolate, S. hominis A9, through full wet-lab characterization (isolation → ID → culture → quantification) and into a Phase 1 RCT in 54 AD adults — twice-daily topical for 7 days reduced S. aureus burden by 99% in 70% of subjects. Every step is wet-lab; AI is essentially absent. What this lineage shows: when the wet-lab is rigorous, you can go to clinic without AI. AI is the compression tool, not the substitute.
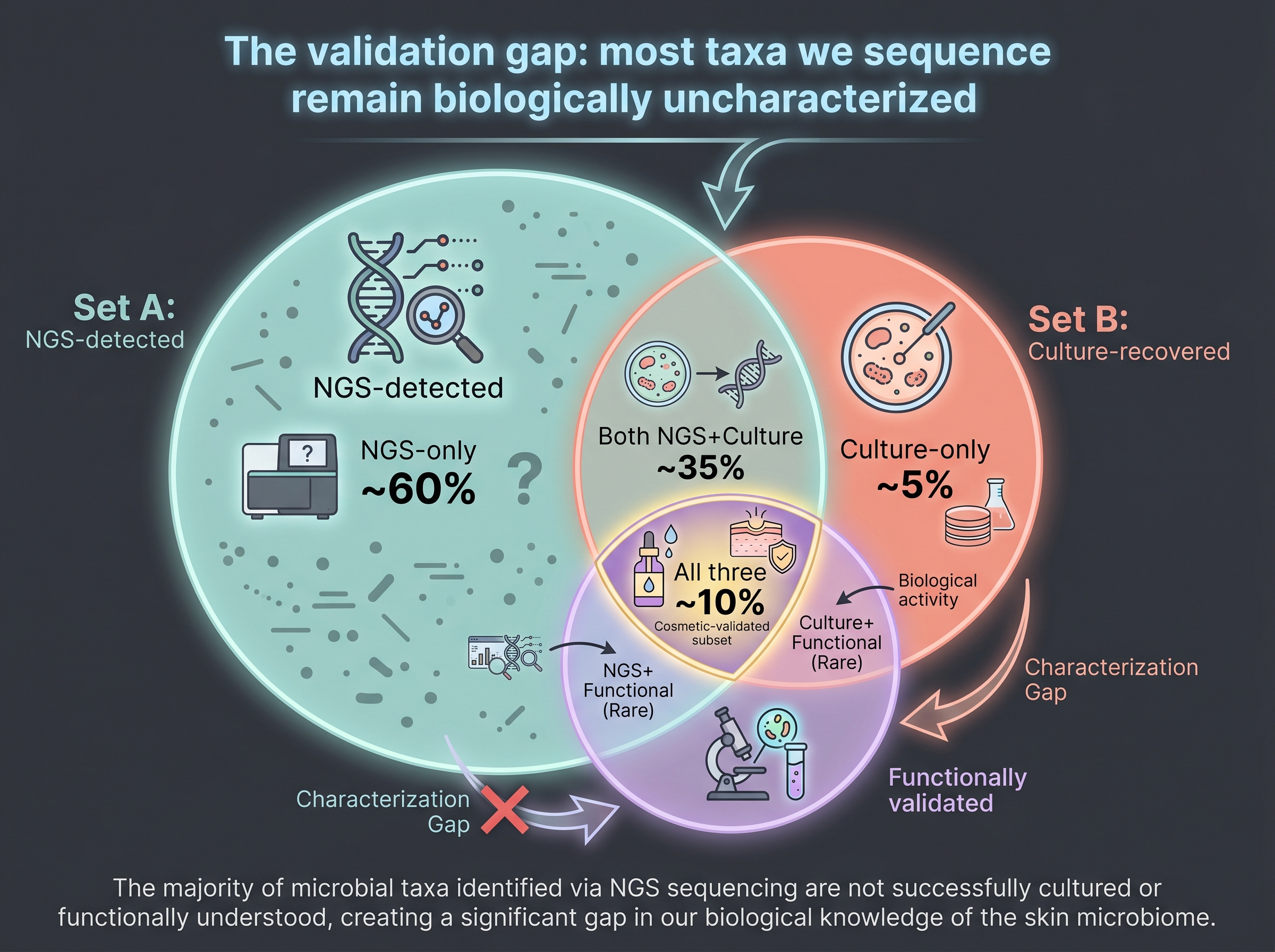
3.7 So — Are Fundamentals Obsolete? Brief A, Answered
Brief A (Chapter 1) asks: "Are isolation, identification, culture, and basic antimicrobial / anti-inflammatory assays now outdated?"
The book's answer is no — they are complementary. Three steps.
- AI compresses search and prediction cost. Narrowing tens of thousands of strain or metabolite candidates to a tractable antimicrobial / anti-inflammatory / anti-aging shortlist now takes one or two orders of magnitude less time than wet-lab-only screening (Chapter 4, Chapter 5).
- Culture and assay validate that prediction. No model proves actual efficacy. An AlphaFold-predicted interaction is a docking score, not a measured in-vitro effect. (Chapter 5) treats this carefully.
- Validated strains and metabolites become the next round's training data. A company that closes this loop accumulates a data asset; one that runs them as siloes restarts every project. This is the (Chapter 7) DBTL primitive.
So "is fundamental research obsolete?" is the wrong framing. The right question is: does our R&D organization bind the AI-prediction layer and the culture-validation layer into one pipeline, or do they sit in separate silos? Only the bound configuration unlocks (Chapter 9) clinical simulation and (Chapter 12) the next-research blueprint.
[3] in Nature Reviews Microbiology explicitly note that "the limited availability of skin-specific reference genomes still constrains metagenomic resolution" — without simultaneous strengthening of culturomics and NGS, that constraint stays. [8] built a foundational reference but covered only 3 skin sites, and as [11] showed, East-Asian cohorts reveal an entirely different catalog. Filling the reference gaps is unfinished work.
3.8 Open Questions — Reproducibility, Cost, Standardization
Open questions across the corpus. The chapter cannot close them; the reader should hold them as we move into Part II.
- How do we control kit and batch effects? Different DNA extraction kits produce statistically different alpha diversities on the same sample. [16] names batch effects as a consensus risk for microbiome ML, but the cosmetic industry has no formal standard analogous to clinical-trial harmonization. (See gaps Gap 8.)
- Ethnic and geographic bias of references. iHSMGC is Han Chinese; HMP is North American. Korean, Japanese, Southeast-Asian, African, and Latin-American skin references are largely empty at cohort scale, and AI efficacy generalization weakens accordingly.
- Cost vs depth. With a fixed budget, increase cohort size? Sample-level depth? Switch to long-read? The cosmetic-endpoint-optimal allocation is not agreed.
- How to accelerate skin culturomics. The Marseille gut culturomics formula was 70+ media + MALDI-TOF. What media, gas mixtures, and identification pipelines does the skin version need? Industry collections accrue privately; no shared protocol exists.
- Absence of a foundation model. No skin-microbiome foundation model exists at ESM3 / DNABERT scale ([9]; cf. [7] ESM3). While data sits inside three companies (Chapter 12), the academic path is narrow.
These five become the entry points to the next chapters: (Chapter 4) shows how AI screening partly sidesteps these limits; (Chapter 12) lays out the data-and-infrastructure blueprint that industry and academia would have to share to close them.
References
- Abubucker, S., Segata, N., Goll, J., et al. (2012). Metabolic reconstruction for metagenomic data and its application to the human microbiome. PLoS Computational Biology.
- Bolyen, E., Rideout, J. R., Dillon, M. R., et al. (2019). Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nature Biotechnology.
- Byrd, A. L., Belkaid, Y., & Segre, J. A. (2018). The human skin microbiome. Nature Reviews Microbiology.
- Callahan, B. J., McMurdie, P. J., Rosen, M. J., Han, A. W., Johnson, A. J. A., & Holmes, S. P. (2016). DADA2: High-resolution sample inference from Illumina amplicon data. Nature Methods.
- Caporaso, J. G., Kuczynski, J., Stombaugh, J., et al. (2010). QIIME allows analysis of high-throughput community sequencing data. Nature Methods.
- Findley, K., Oh, J., Yang, J., et al. (2013). Topographic diversity of fungal and bacterial communities in human skin. Nature.
- Hayes, T., et al. (2025). Simulating 500 million years of evolution with a language model (ESM3). Science. [Hayes et al., 2025]
- Human Microbiome Project Consortium (2012). Structure, function and diversity of the healthy human microbiome. Nature. [HMP Consortium, 2012]
- Wang, X.-W., Wang, T., & Liu, Y.-Y. (2024). Artificial Intelligence for Microbiology and Microbiome Research. arXiv. [Wang et al., 2024]
- Lagier, J.-C., Edouard, S., Pagnier, I., Mediannikov, O., Drancourt, M., & Raoult, D. (2018). Culturing the human microbiota and culturomics. Nature Reviews Microbiology.
- Li, Z., Xia, J., Jiang, L., et al. (2021). Characterization of the human skin resistome and identification of two microbiota cutotypes (iHSMGC). Microbiome.
- Nakatsuji, T., Chen, T. H., Narala, S., et al. (2017). Antimicrobials from human skin commensal bacteria protect against Staphylococcus aureus and are deficient in atopic dermatitis. Science Translational Medicine.
- Nakatsuji, T., Hata, T. R., Tong, Y., et al. (2021). Development of a human skin commensal microbe for bacteriotherapy of atopic dermatitis and use in a phase 1 randomized clinical trial. Nature Medicine.
- Oh, J., Byrd, A. L., Deming, C., Conlan, S., NISC Comparative Sequencing Program, Kong, H. H., & Segre, J. A. (2014). Biogeography and individuality shape function in the human skin metagenome. Nature.
- Oh, J., Byrd, A. L., Park, M., NISC Comparative Sequencing Program, Kong, H. H., & Segre, J. A. (2016). Temporal Stability of the Human Skin Microbiome30399-3). Cell.
- Papoutsoglou, G., et al. (2023). Machine learning approaches in microbiome research: challenges and best practices. Frontiers in Microbiology. [Papoutsoglou et al., 2023]
- Quast, C., Pruesse, E., Yilmaz, P., Gerken, J., Schweer, T., Yarza, P., Peplies, J., & Glöckner, F. O. (2013). The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Research.
- Truong, D. T., Franzosa, E. A., Tickle, T. L., Scholz, M., Weingart, G., Pasolli, E., Tett, A., Huttenhower, C., & Segata, N. (2015). MetaPhlAn2 for enhanced metagenomic taxonomic profiling. Nature Methods.