Chapter 2: The Skin Microbiome — A Map of Sites, Species, and Homeostasis
2.1 Why "the skin microbiome" is a useful concept
Human skin is a complex environment — roughly 1.8 m² of surface area folded into pores, wrinkles, glands, and follicles. Living on that surface is a microbial population comparable in cell count to the host's own. These microbes are not contamination to be wiped away; they are a symbiotic ecosystem that tunes immune signaling and barrier function. Where first-generation fermentation cosmetics (Chapter 1) asked "which strain produces which active fermentate?", second-generation microbiome cosmetics shifted the question itself: "Which microbial community produces which skin state, and how can we modulate it?"
The shift has roots in the late 1990s, but the decisive year was 2009. That year two Science papers delivered the same message from different angles — skin is populated by site-specific communities, and those communities depend more on body site than on individual identity [7] [4]. The two findings together became the starting coordinates of every skin-microbiome cosmetic R&D effort since.
The three quantitative anchors of this chapter 1. Site beats individual: your palm community looks more like someone else's palm than like your own forehead [4]. 2. Inter-individual variation is large but temporal stability is high: at sebaceous sites, more than 80% of strains in a person persist over 1–2 years [18]. 3. Species is too coarse — strain decides phenotype: acne patients and healthy individuals carry similar total Cutibacterium acnes abundance, but different strain distributions [6].
The purpose of this chapter is to draw the map the rest of the book repeatedly refers to. Site ecology (2.2) → core taxa (2.3) → inter-individual variation and ethnic differences (2.4) → an anatomy of the word "dysbiosis" (2.5) → correlation and causation with disease (2.6) → open questions (2.7). To understand what AI models in Chapters 4–6 actually learn — and where they fail — you need this map first.
2.2 The site framework — sebaceous, moist, dry
The hardest empirical rule in skin microbiome work is that site dominates everything. [7] sequenced 16S rRNA from 20 body sites in 10 healthy adults and classified skin by physical-physiological habitat into three categories.
Sebaceous (lipid-rich) sites — forehead, glabella, nose, upper back. The lipid-rich environment is occupied by Cutibacterium acnes (formerly Propionibacterium acnes), which thrives on sebum hydrolysis. Within pilosebaceous units, C. acnes commonly accounts for more than 70% of relative abundance [2].
Moist sites — axilla, antecubital and popliteal fossae, toe webs. Staphylococcus and Corynebacterium dominate, favored by high humidity and salinity.
Dry sites — forearms, palms, legs. These host the most diverse communities — about 44 species on average, versus 19 at the retroauricular crease (the least diverse site) [7] — with Proteobacteria and Flavobacteriales more prominent than at the lipid-rich or moist sites.
The same year, [4] generalized this site framework across the entire body. Pooling skin, oral, gut, and urogenital samples, they showed that the variable most strongly structuring microbial community composition was site > individual > time. Your forehead microbiome looks more like a stranger's forehead than your own forearm.
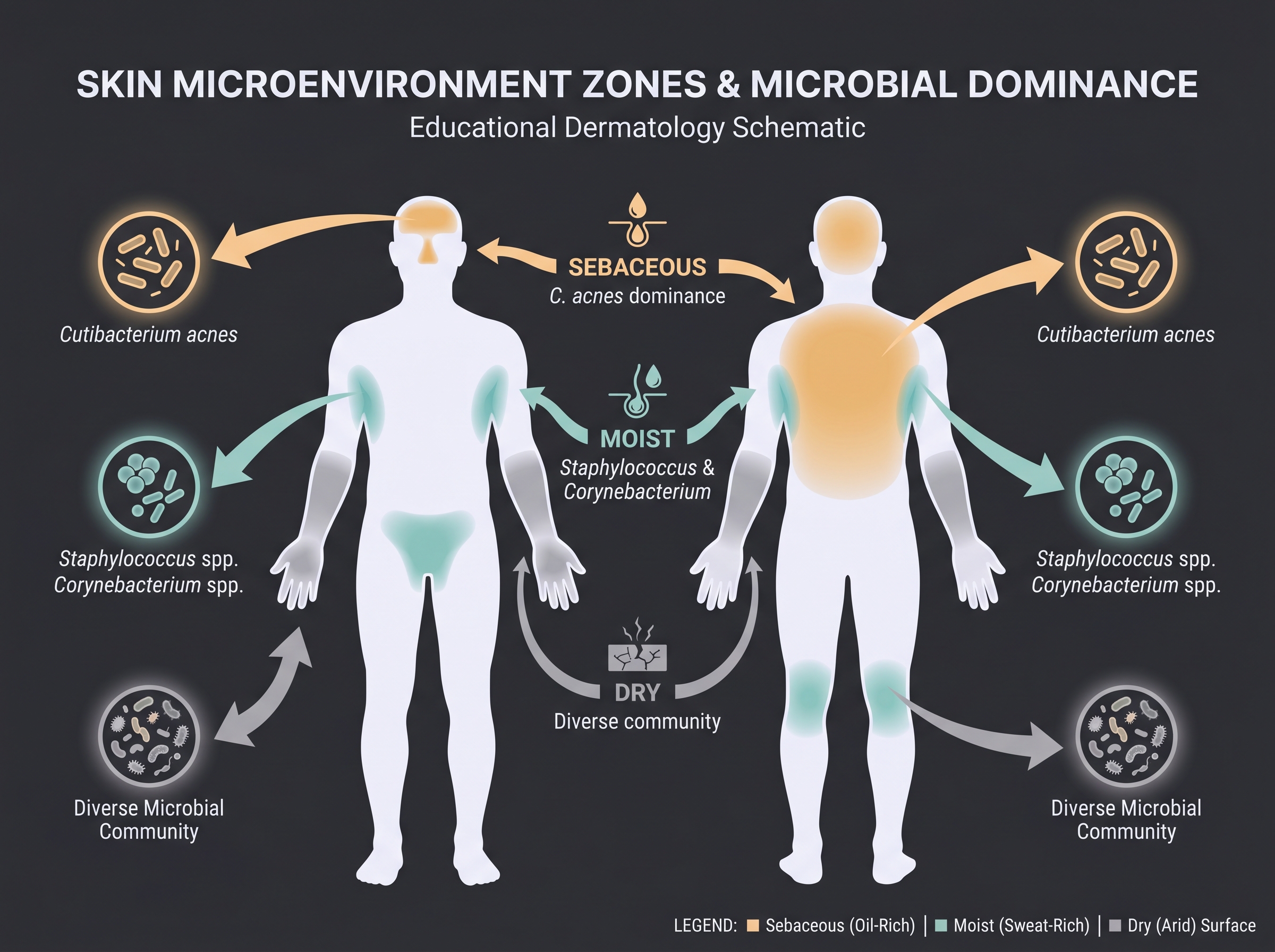
The framework is simple but powerful. For cosmetic R&D, "which site does this product touch?" is already 80% of "which microbial community does it interact with?" The community a facial serum encounters is almost entirely different from the one a deodorant meets. Even when evaluating microbiome-friendly cleansers, lumping the T-zone and U-zone into one endpoint dilutes the signal.
But the three-bucket classification has a limitation worth naming: it discretizes a continuous variable. Sebum production varies smoothly across regions, and an individual's sebum level depends on genetics, hormones, and seasonality. One difficulty later AI models (Chapter 6) repeatedly face is the trade-off between learning the labels sebaceous/moist/dry and learning the continuous measurements sebum, transepidermal water loss (TEWL), pH. Both work; the latter is closer to the underlying biology.
2.3 Core taxa — Cutibacterium acnes, Staphylococcus epidermidis, Malassezia
Hundreds of species inhabit skin, but nearly every cosmetic R&D discussion eventually converges on three taxa (or genera). They matter not just for their abundance but because they are functionally decisive.
2.3.1 Cutibacterium acnes — the two faces of the sebaceous gland
Renamed in 2016 from Propionibacterium acnes to Cutibacterium acnes [2], this organism is both the most common and the most misunderstood resident of sebaceous skin. For three decades the cosmetic and dermatology industries built antimicrobial strategies on the simple equation "acne = P. acnes." That premise was never tested at strain resolution.
When [6] analyzed pilosebaceous units from 49 acne patients and 52 healthy controls at strain resolution, the result was startling — the two groups carried statistically indistinguishable total C. acnes abundance. The difference was in strain composition. RT4 and RT5 ribotypes were enriched in acne skin, RT6 in healthy skin. C. acnes is not the enemy; it is a dual-natured resident — some strains protective, others pathogenic.
This finding inverted cosmetic strategy. "Kill C. acnes" approaches can remove protective strains together with pathogenic ones, leaving the long-term ecosystem less stable. New strategies have emerged instead — strain-selective phage cocktails, engineered strains overexpressing RoxP (the natural antioxidant some C. acnes phylotypes secrete), and ribotype-stratified diagnostics. (We return to this industry branch in Chapter 10.)
2.3.2 Staphylococcus epidermidis — the protective commensal
S. epidermidis can account for up to 90% of aerobic resident skin bacteria, making it the most prevalent commensal [2]. Until the 1990s clinical microbiology dismissed it as "hospital-infection contaminant." [13] inverted that assessment at mechanism resolution.
The circuit they described: lipoteichoic acid (LTA) from the S. epidermidis cell wall binds keratinocyte Toll-like receptor 2 (TLR2). The resulting TLR2 signal suppresses TLR3 signaling within the same cell. TLR3 responds to self-RNA released during injury and, unchecked, drives excessive inflammation. So when S. epidermidis is present on the surface, inflammatory responses to barrier damage are automatically dampened. This was the first molecular-level mechanism for the 30-year-old claim that "probiotics are good for skin."
[15] then used germ-free mice to show that S. epidermidis is necessary for IL-1-dependent T-cell education in skin, and the Gallo group [16] demonstrated that coagulase-negative staphylococci (CoNS) such as S. hominis A9 secrete antimicrobial peptides that block S. aureus colonization. S. epidermidis and its relatives are not "harmless co-inhabitants" — they are part of the local immune system.
A subtle trap remains. Some S. epidermidis strains form biofilms on catheters and prosthetic joints, causing nosocomial infections. Like S. aureus, S. epidermidis shows large strain-level phenotypic variance. Which strain produced which postbiotic or equivalent — and how it was isolated and purified — matters far more than marketing copy admits.
2.3.3 Malassezia — the fungal heavyweight
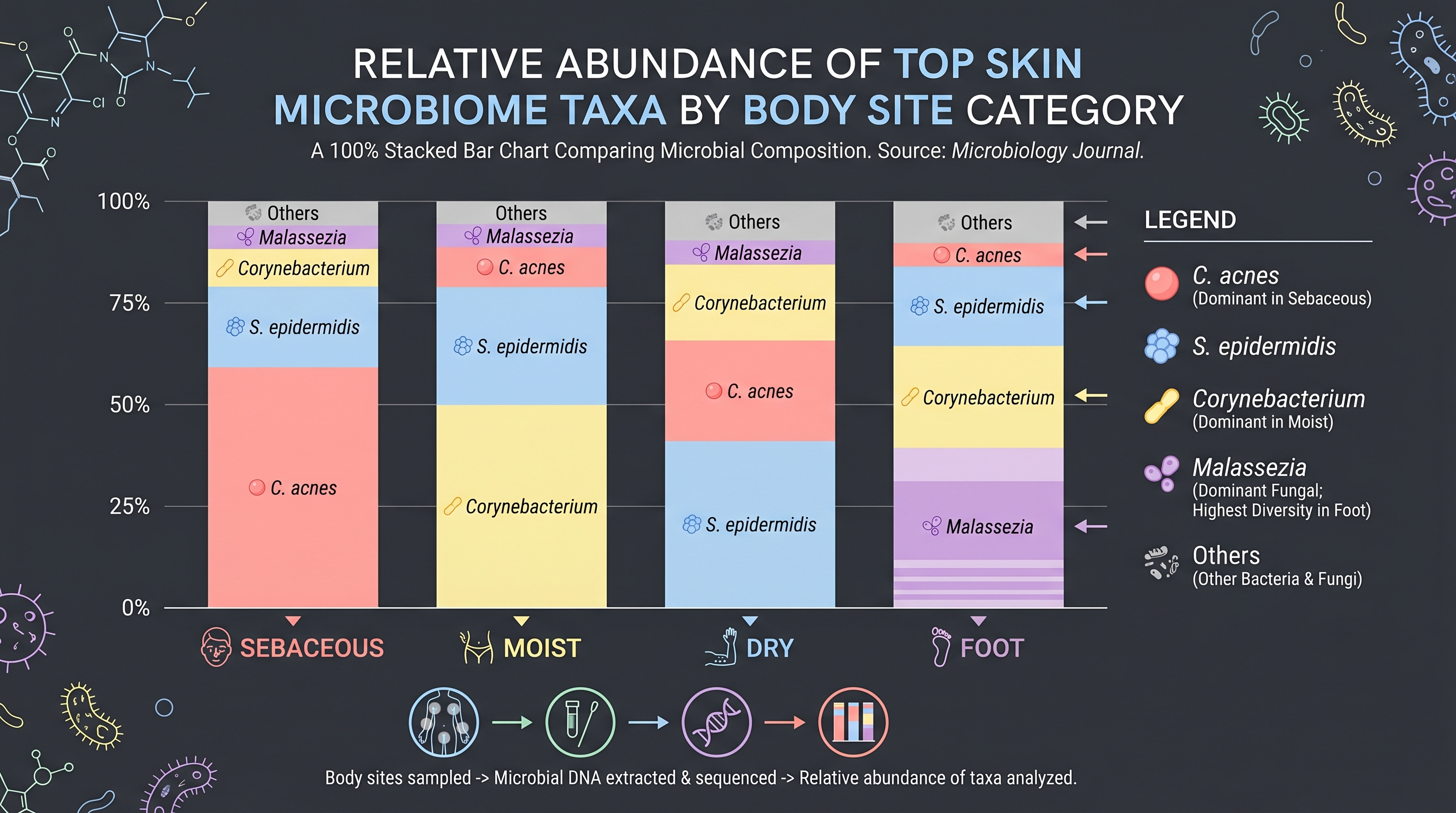
[5] disrupted the bacterial-only framing of early microbiome work by simultaneously sequencing fungal ITS1 and bacterial 16S. They reported that more than 90% of skin fungal communities belong to a single genus, Malassezia. At 11 head/arm/trunk sites Malassezia accounted for over 80% of fungal reads, while foot sites — toes, plantar heel, toenail — were 2–5× more diverse and included Aspergillus, Cryptococcus, and Rhodotorula.
Malassezia is lipid-dependent. Unable to synthesize fatty acids itself, it hydrolyzes sebum triglycerides for survival. Lipases released in that process generate free fatty acids — particularly oleic acid — that irritate some individuals' skin and underlie dandruff and seborrheic dermatitis [19]. This is why the active ingredients in dandruff shampoos (zinc pyrithione, ketoconazole, selenium sulfide) are all antifungals.
But Malassezia is also a normal resident. As with C. acnes, the question is not "present or absent" but "which species and how much." M. restricta and M. globosa are the two most common species, but their ratio and metabolic activity shape clinical phenotypes from seborrheic dermatitis to atopic dermatitis to Malassezia folliculitis. Modeling bacteria and fungi as a single multi-kingdom ecosystem is one of the central problems of Chapter 6.
2.4 Inter-individual variation — twins, ethnicity, and East Asia's "hidden genes"
If site explains 80% of community composition, what explains the rest? The answer is messy. Genetics, age, hormones, geography, lifestyle, and cosmetic use history all contribute. But the single most striking finding came from [14].
[14] collected shotgun metagenomic data from the skin of 822 Han Chinese individuals and combined it with 538 prior North American samples to build the integrated Human Skin Microbial Gene Catalog (iHSMGC). They cataloged 10.93 million microbial genes — and about 4.88 million, or 44.6%, were entirely new relative to existing catalogs. Even more striking, Moraxella osloensis was the second most abundant species in the Chinese cohort but essentially absent from the North American one. The authors defined two "cutotypes" — a Moraxella osloensis type and a C. acnes type — and explained the split with a host-dependent trophic-chain hypothesis.
The implication is deep. Every ML model trained only on North American or European cohorts has never seen roughly 45% of the microbial genes relevant to East Asian consumers. This is the strategic motivation for Korean, Chinese, and Japanese firms building their own metagenomic cohorts. (How that asymmetry plays out in Korean industry and academia is the subject of Chapter 10.)
Twin studies add another angle. Monozygotic twins share more similar skin microbiomes than dizygotic twins, but the genetic effect is small compared with site or age. Family and roommate effects (shared home environments) are also statistically significant. The picture that emerges is of a microbiome shaped less by personal identity and more by person × environment.
Finally, temporal stability. [18] resampled 12 of the original HMP subjects 1–2 years later and showed that more than 80% of strains persisted at strain resolution. Sebaceous sites were most stable, feet least. The implication cuts both ways — first, the microbiome is stable enough to call "yours"; second, it is hard to durably shift with topical cosmetics. The "restore your microbiome" marketing line often collides with this stability.
2.5 Anatomizing the word "dysbiosis"
The headline word of microbiome cosmetic marketing is "dysbiosis." Advertising claims that dysbiosis causes acne, atopic dermatitis, and aging, and that the product "restores balance." In the academic literature, dysbiosis is surprisingly slippery.
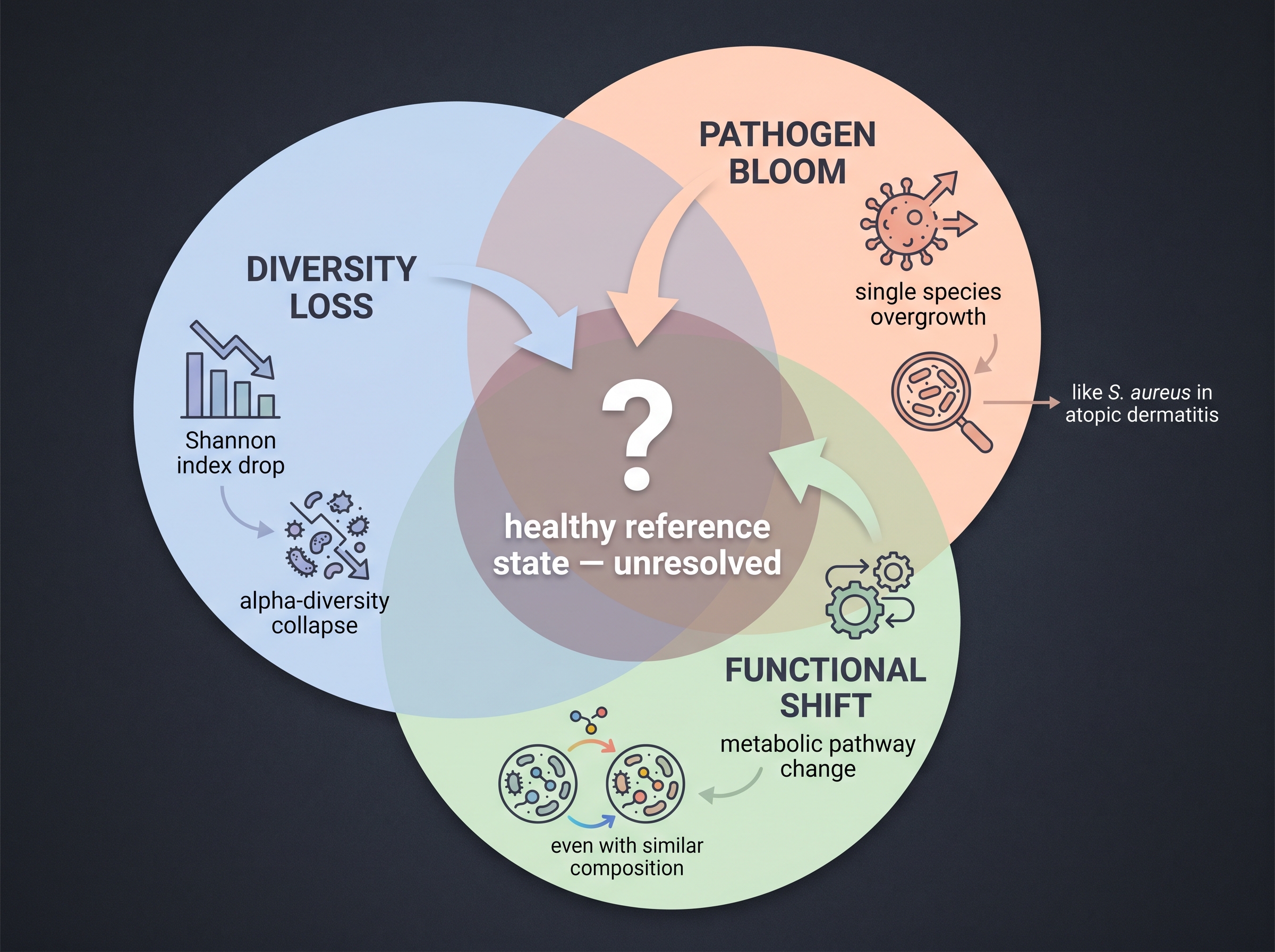
There are at least three operational definitions [2]:
- Diversity loss — Shannon diversity below the healthy-cohort mean.
- Overgrowth of pathogenic taxa — S. aureus replacing S. epidermidis, for instance.
- Functional dysbiosis — abnormal gene or metabolic profiles regardless of taxonomic abundance.
The three definitions can disagree on the same dataset. In the atopic dermatitis cohort of [11], S. aureus surges during flare while Shannon diversity simultaneously falls. After treatment, diversity recovers but clinical improvement begins before S. aureus fully clears. The statement "dysbiosis improved" picks a different timepoint depending on which definition you mean.
The harder problem is the reference state. There is no agreed-upon standard for a "healthy" skin microbiome. The Human Microbiome Project defined one for North American adults, iHSMGC added a Han Chinese reference, but no standard cohort exists that controls jointly for age, season, lifestyle, and prior cosmetic use. A cosmetic clinical study that names "dysbiosis improvement" as an endpoint should specify upfront which quantitative criterion defines dysbiosis and which defines its reversal. Published cosmetic trials with explicit prespecification of this kind remain rare [1].
[3] put the criticism more bluntly — dysbiosis can refer to "any statistical deviation in microbial community composition" and therefore cannot, by itself, support a causal claim. This book uses the term with that caveat in mind, naming the operational definition each time and separating it from the marketing usage.
2.6 Connection to disease — the line between correlation and causation
Even with the slipperiness of "dysbiosis," the association between the microbiome and skin disease is well established. The interpretation of that association is where the difficulty lies.
Acne. The strongest signal is strain-level shifts in C. acnes [6]. RT4 and RT5 ribotypes are enriched in lesional skin and carry plausible virulence factors such as hyaluronate lyase and CAMP factor. But whether the strain shift causes acne or acne creates the niche that selects the strain shift is still unresolved. Causation runs through the interaction of pilosebaceous-unit microenvironment (sebum composition, oxygen tension) and strain fitness, and does not collapse to a single-direction arrow.
Atopic dermatitis (AD). The first quantitative model came from [11] — at flare, S. aureus surges and protective commensals like S. epidermidis decline. Whether the microbial shift is cause or consequence of AD remains actively debated. Host factors such as filaggrin loss-of-function variants weaken the epidermal barrier, which lets S. aureus establish more easily. From that angle, microbial change is a marker of host defect, not a driver. But [16] showed clinical improvement after direct microbial intervention (transplanting protective CoNS strains) — suggesting the microbiome is at least partially a modifiable causal factor.
The two views are not exclusive. The most credible model is a multifactorial circuit of host × microbiome × environment. AD is the state in which that circuit is trapped in a self-amplifying loop: filaggrin deficiency → barrier damage → S. aureus expansion → further barrier damage. A microbiome cosmetic can perturb one node (the microbial one); it cannot perturb others (host genotype).
Rosacea. Serum reactivity to Bacillus oleronius proteins carried by Demodex mites is 82.6% in rosacea patients versus 26.9% in controls [12]. Strong association, ambiguous causation — does the rosacea skin environment permit Demodex expansion, or does Demodex drive rosacea? Not yet settled.
Seborrheic dermatitis. Strongly linked to Malassezia abundance, especially M. restricta [19]. Here the causal evidence is comparatively strong — antifungal therapy improves the clinical picture.
What these conditions share is that microbial shifts are consistently observed, but whether they are cause or consequence varies by disease and is often bidirectional. For cosmetic R&D, this distinction is decisive. A cosmetic that targets the consequence offers short-term symptomatic relief; a cosmetic that targets the cause — when that is possible — offers more fundamental improvement. Most marketed microbiome cosmetics claim "restored balance" without specifying which.
2.7 Open Questions — what this chapter does not resolve
The map is drawn, but blank patches remain. The rest of the book returns to three questions over and over.
Q1. How should dysbiosis be operationalized? Shannon diversity, taxon ratios, functional-profile distance — which single metric correlates best with clinical endpoints? And can it be measured reliably enough to use as a regulatory or claims-grade outcome? Chapter 3 (NGS methodology) and Chapter 9 (clinical endpoint design) return to this question.
Q2. How should inter-individual variation be interpreted — noise or signal? The "your microbiome is yours" identity view is the marketing foundation of personalized cosmetics. But how much of that variation is explained by measurement noise, environmental variance, or detection limits? Chapter 6 (interaction modeling) and Chapter 9 (clinical efficacy validation) attempt answers.
Q3. How can temporal stability and cosmetic efficacy coexist? [18] showed >80% strain-level stability over 1–2 years. If a cosmetic "restores" the microbiome, is that restoration permanent or transient? Does the user revert when use stops? This question sits at the core of the long-term value proposition of microbiome cosmetics. Chapter 9 and Chapter 12 confront it directly.
These three questions reappear, in progressively more concrete forms, throughout the rest of the book. AI (Part II) tries new answers and runs into new versions of the same limitations. Synthetic biology and clinical validation (Part III) bring experimental pressure to force answers. Industry (Part IV) shows what it looks like to make decisions while the answers are still missing.
The map is a beginning, not an end. In the next chapter (Chapter 3) we turn to the methods that drew it — NGS, metagenomics, and the resurgent culturomics that complements them.
References
- Atallah, C., Charcosset, C., Greige-Gerges, H. (2025). Bioengineered Skin Microbiome — A Critical Review on Skin Microbiome Modulation Strategies. Cosmetics, 12(5), 205. DOI: 10.3390/cosmetics12050205.
- Byrd, A. L., Belkaid, Y., Segre, J. A. (2018). The human skin microbiome. Nature Reviews Microbiology, 16(3), 143–155. DOI: 10.1038/nrmicro.2017.157.
- Carmona-Cruz, S., Orozco-Covarrubias, L., Saez-de-Ocariz, M. (2022). The Human Skin Microbiome in Selected Cutaneous Diseases. Frontiers in Cellular and Infection Microbiology, 12, 834135. DOI: 10.3389/fcimb.2022.834135. [Carmona-Cruz et al., 2022]
- Costello, E. K., Lauber, C. L., Hamady, M., Fierer, N., Gordon, J. I., Knight, R. (2009). Bacterial Community Variation in Human Body Habitats Across Space and Time. Science, 326(5960), 1694–1697. DOI: 10.1126/science.1177486.
- Findley, K., Oh, J., Yang, J., Conlan, S., Deming, C., Meyer, J. A., et al. (2013). Topographic diversity of fungal and bacterial communities in human skin. Nature, 498(7454), 367–370. DOI: 10.1038/nature12171.
- Fitz-Gibbon, S., Tomida, S., Chiu, B.-H., Nguyen, L., Du, C., Liu, M., et al. (2013). Propionibacterium acnes strain populations in the human skin microbiome associated with acne35970-1/fulltext). Journal of Investigative Dermatology, 133(9), 2152–2160. DOI: 10.1038/jid.2013.21.
- Grice, E. A., Kong, H. H., Conlan, S., Deming, C. B., Davis, J., Young, A. C., et al. (2009). Topographical and Temporal Diversity of the Human Skin Microbiome. Science, 324(5931), 1190–1192. DOI: 10.1126/science.1171700.
- Grice, E. A., Segre, J. A. (2011). The skin microbiome. Nature Reviews Microbiology, 9(4), 244–253. DOI: 10.1038/nrmicro2537. [Grice & Segre, 2011]
- Haykal, D., Flament, F., Amar, D., Cartier, H., Kourosh, A. S., Lee, S., Rowland-Payne, C. (2025). Cosmetogenomics unveiled: a systematic review of AI, genomics, and the future of personalized skincare. Frontiers in Artificial Intelligence, 8, 1660356. DOI: 10.3389/frai.2025.1660356.
- Human Microbiome Project Consortium (2012). Structure, function and diversity of the healthy human microbiome. Nature, 486(7402), 207–214. DOI: 10.1038/nature11234. [HMP Consortium, 2012]
- Kong, H. H., Oh, J., Deming, C., Conlan, S., Grice, E. A., Beatson, M. A., et al. (2012). Temporal shifts in the skin microbiome associated with disease flares and treatment in children with atopic dermatitis. Genome Research, 22(5), 850–859. DOI: 10.1101/gr.131029.111.
- Lacey, S., Kavanagh, S., Powell, S. C. (2014). Correlation between serum reactivity to Demodex-associated Bacillus oleronius proteins, and altered sebum levels and Demodex populations in erythematotelangiectatic rosacea patients. Journal of Medical Microbiology, 63(11), 1456–1462. DOI: 10.1099/jmm.0.073262-0.
- Lai, Y., Di Nardo, A., Nakatsuji, T., Leichtle, A., Yang, Y., Cogen, A. L., et al. (2009). Commensal bacteria regulate Toll-like receptor 3-dependent inflammation after skin injury. Nature Medicine, 15(12), 1377–1382. DOI: 10.1038/nm.2062.
- Li, Z., Xia, J., Jiang, L., Tan, Y., An, Y., Zhu, X., et al. (2021). Characterization of the human skin resistome and identification of two microbiota cutotypes. Microbiome, 9(1), 47. DOI: 10.1186/s40168-020-00995-7. [Li et al., 2021]
- Naik, S., Bouladoux, N., Wilhelm, C., Molloy, M. J., Salcedo, R., Kastenmuller, W., et al. (2012). Compartmentalized Control of Skin Immunity by Resident Commensals. Science, 337(6098), 1115–1119. DOI: 10.1126/science.1225152.
- Nakatsuji, T., Chen, T. H., Narala, S., Chun, K. A., Two, A. M., Yun, T., et al. (2017). Antimicrobials from human skin commensal bacteria protect against Staphylococcus aureus and are deficient in atopic dermatitis. Science Translational Medicine, 9(378), eaah4680. DOI: 10.1126/scitranslmed.aah4680.
- Oh, J., Byrd, A. L., Deming, C., Conlan, S., NISC Comparative Sequencing Program, Kong, H. H., Segre, J. A. (2014). Biogeography and individuality shape function in the human skin metagenome. Nature, 514(7520), 59–64. DOI: 10.1038/nature13786. [Oh et al., 2014]
- Oh, J., Byrd, A. L., Park, M., NISC Comparative Sequencing Program, Kong, H. H., Segre, J. A. (2016). Temporal Stability of the Human Skin Microbiome30399-3). Cell, 165(4), 854–866. DOI: 10.1016/j.cell.2016.04.008.
- Smith, A. (2025). Skin Microbiome in Health and Disease — A Comprehensive Review. Microorganisms, 13(1), Jan 2025. [Smith, 2025]