Chapter 1: Microbes in Cosmetics — A History of Fermentation and Its Limits
1.1 Why this chapter — the word "fermentation" has changed meaning twice
At a cosmetics counter, the word "ferment" has changed meaning twice — once in the 1980s, and once again in the mid-2010s. The first shift began with a Japanese technician noticing that brewers' hands, kept in contact with a yeast-fermented liquid, looked unusually smooth. That observation became SK-II Pitera, launched in 1980 as a single-strain fermentation filtrate. The second shift happened when the molecular mechanism of that filtrate was finally described at the molecular level by [17] in Clinical and Experimental Dermatology — thirty-five years after commercial launch.
That thirty-five-year gap is the subject of this chapter. To understand what microbiome cosmetics actually are and why AI is being called an inflection point, we have to be honest about what the first generation of fermentation cosmetics did — and what it did without knowing why. A product category that survived for three decades on marketing language is now being rewritten under the data-driven instruments of metagenomics, AI, and synthetic biology. The rewrite is the starting point of the book.
This chapter assumes a smart non-expert reader — a cosmetic marketer, journalist, or investor — and explicitly avoids molecular-biology prerequisites. The concepts and open questions that subsequent chapters (Chapter 3, Chapter 5) will lean on are introduced here so that the reader does not have to retrace them later.
The three quantitative anchors of this chapter
- 35 years — the gap between commercial launch (SK-II Pitera, 1980) and the first molecular mechanism paper ([17]).
- ~1,000-fold — the drop in human-genome sequencing cost from roughly $10M in 2005 to roughly $1,000 in 2014 (US NHGRI Cost-per-Genome data). Without this collapse, the skin microbiome atlas could not have existed.
- 205+ entries — the cumulative 2025 indexing of microbiome/bioengineered cosmetic papers and reviews in a single MDPI journal, Cosmetics (e.g., [1] as the closing issue of the volume).
1.2 First-generation fermentation cosmetics — what they did, and what was an accident
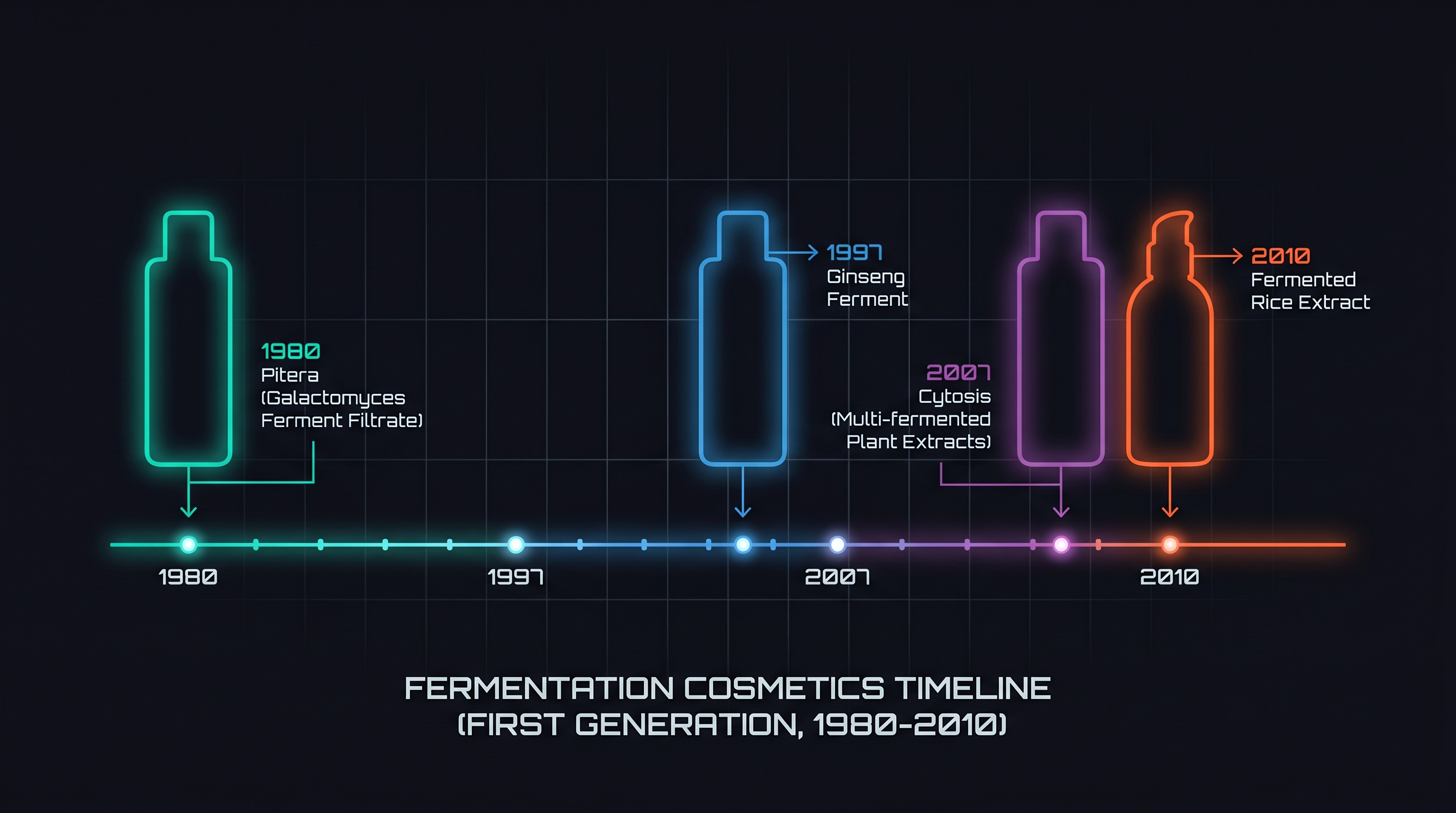
First-generation fermentation cosmetics rested on three empirical observations.
Observation 1 — Ferment filtrates appeared to have anti-aging and soothing effects. SK-II Pitera began as a filtered liquid from a sake-brewing process driven by Galactomyces (or Pichia) yeast. Through the 1990s in Japan and Korea, a wave of "Korean fermentation cosmetics" emerged — botanical donguibogam-derived recipes fermented by yeast or lactic acid bacteria. By the early 2010s, brands such as Sulwhasoo, The History of Whoo, SUM37, and Hera had built fermentation lines into multi-billion-dollar revenue categories.
Observation 2 — Ferment filtrates of Lactobacillus, Bifida, and Saccharomyces showed clinically measurable hydration and barrier signals. These signals were partially confirmed through in vitro keratinocyte assays and 12-week placebo-controlled trials in the early-to-mid 2000s. But almost every such study was framed as single-ferment vs vehicle — none answered "which molecule in the filtrate binds which receptor to switch which gene."
Observation 3 — The fermentate is safer than the live organism. Putting live bacteria into cosmetics multiplies preservation, stability, and regulatory burdens, so essentially every first-generation fermentation product was a postbiotic — a killed-cell or cell-free filtrate preparation. This was driven by operational constraints, not by molecular insight.
In one line: first-generation fermentation cosmetics reached empirical efficacy without reaching mechanistic explanation. [12] make the same diagnosis at the probiotic-industry level — the mechanism-deficit was not a cosmetic-specific failure, but a structural condition of the whole probiotic field.
1.2.1 Galactomyces — a thirty-five-year mechanism gap, finally filled
[17] was the first paper to describe the molecular mechanism of a first-generation fermentation product at depth. The authors showed that Galactomyces ferment filtrate (GFF) activates the aryl hydrocarbon receptor (AHR) in keratinocytes, and that AHR activation restores filaggrin expression suppressed by Th2 cytokines (IL-4, IL-13). AHR is the nuclear receptor that senses tryptophan-derived and microbiome-derived metabolites; filaggrin is the key skin-barrier protein.
Seven years later, [7] added a second molecular axis: the same Galactomyces extract switches on the NRF2 (nuclear factor erythroid 2-related factor 2) antioxidant axis (HO-1, NQO1) and dampens NF-κB-mediated inflammation. AHR + NRF2 — when both axes are jointly explained, the brewers' smooth hands of the 1980s finally have a molecular account.
Two implications follow. First, the empirical efficacy was not a fiction — only the explanation was late. Second, the thirty-five-year delay was a function of when AHR became a mainstream tool in skin biology (late 1990s onward), and the modern tool for compressing that timeline is the kind of protein-structure prediction we will meet in Chapter 5 (AlphaFold, ESMFold). If the mechanism of a fermentation cosmetic took thirty-five years to fill in, could a candidate molecule from the AI era take thirty-five weeks? That question frames the whole of Part II.
1.2.2 The asymmetry between marketing language and scientific language
The second legacy of first-generation fermentation cosmetics is a language asymmetry. Advertising lived for three decades on slogans — "one hundred efficacies," "imperial recipe," "100 million active yeast cells" — while the peer-reviewed literature on the same products fit on the fingers of one hand. [1] make this explicit in their 2025 review: "Most cited efficacy claims for bioengineered actives rely on company press releases, not peer-reviewed clinical trials."
This asymmetry is not just an ethical problem; it is a structural obstacle to the transition into data-driven second-generation cosmetics. If the vague efficacy language of the first generation is carried over, the precise endpoints made possible by metagenomics and AI (strain-level collagen-synthesis induction, site-specific dysbiosis recovery rates) will be diluted into the same fuzz. Chapter 8 (AI formulation) and Chapter 12 (research blueprint) address this directly.
1.3 The sequencing-cost curve — the technical inflection that ended the first generation
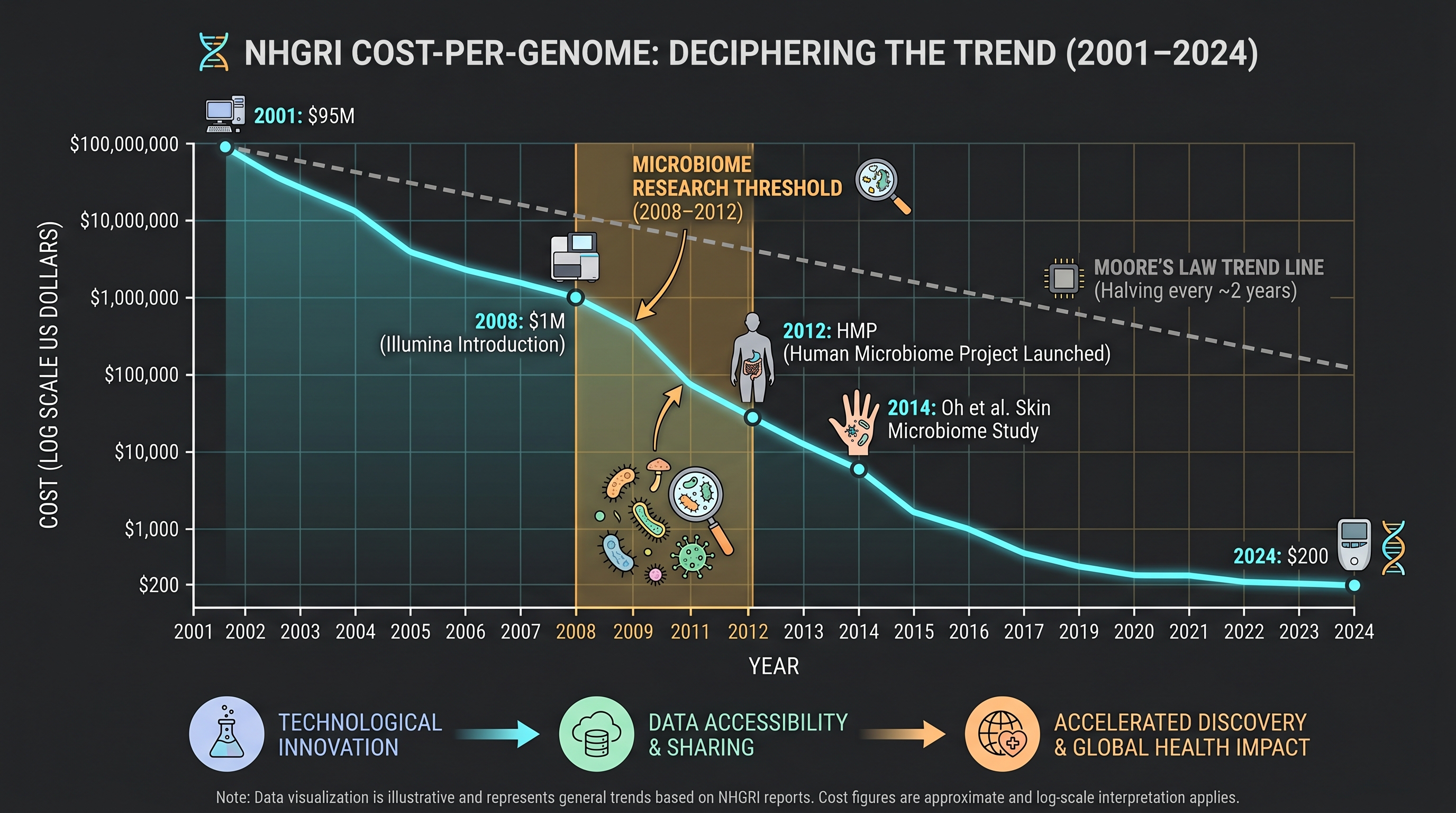
The biggest external constraint that kept first-generation fermentation cosmetics on a single-strain, single-filtrate footing was sequencing cost. In 2001, when the Human Genome Project was declared complete, sequencing one human genome cost about $100M. By 2005, with next-generation sequencing (NGS — particularly 454/Roche pyrosequencing), cost dropped to about $10M. By 2014, with Illumina HiSeq X Ten, cost reached about $1,000 (US NIH National Human Genome Research Institute Cost-per-Genome tracking data).
Translated into the microbiome frame: for first-generation cosmetic R&D planners, seeing what microbes lived across a person's skin was not even an economic option. A single 16S rRNA amplicon sweep of one body site cost thousands of dollars in 2008; shotgun metagenomics on even a single-digit patient cohort was a luxury. Only after the cost curve collapsed did "look at the whole skin ecosystem" become an executable idea.
1.4 Three papers that opened the microbiome era (2009–2012)
Within four years of the cost-curve collapse, the field of human microbiome research acquired its three foundational papers.
[2] — Science. The Rob Knight group sampled 27 body sites in 9 healthy adults, repeated over time, using 16S rRNA sequencing. The conclusion was simple but powerful: body site is a stronger determinant of community structure than inter-individual variation. The palm and the mouth of the same person are more different from each other than two palms across people. This was the first formal answer to why first-generation "put a strain in a jar" thinking only partially worked — different sites are different ecosystems, and the same microbe behaves differently in different ecosystems.
[5] — Science. In the same year, Julia Segre's group published the canonical paper on the skin microbiome atlas. Twenty body sites were grouped into three microenvironments — sebaceous (oily), moist, and dry — and each was shown to have distinct dominant taxa: Propionibacterium (now Cutibacterium) in sebaceous sites, Staphylococcus and Corynebacterium in moist sites, and a more diverse community in dry sites. First-generation cosmetics had divided products into "face" vs "body" by marketing logic — by chance, that division partly aligned with these microenvironments. After 2009, the division could be defended with data.
[8] — Nature. The $170M NIH Human Microbiome Project released its first reference dataset after five years of effort — 242 healthy adults, 15–18 body sites per subject, 16S rRNA plus shotgun metagenomics. The data did not flow directly into cosmetic product pipelines, but it provided what every subsequent disease-or-dysbiosis study needed: a population-scale baseline of what "healthy" looks like. You cannot define dysbiosis without first defining the norm. HMP defined the norm.
Together, these three papers complete a single move. [6], in a Nature Reviews Microbiology synthesis, frame the transition: in the five-year window after these papers, skin microbiome research went from a topic without instruments to a discipline with instruments. The analogy is the microbiology of before and after the microscope.
1.4.1 Fungi, strain resolution, temporal stability — the supporting scaffolding (2013–2016)
Three pieces of supporting scaffolding were laid down right after the founding papers.
[3] in Nature applied ITS (internal transcribed spacer) sequencing to map the fungal community on skin — most importantly the site-by-site distribution of the genus Malassezia. Fermentation cosmetics had handled yeast for thirty years, but the yeast ecosystem that actually lives on human skin was effectively invisible until this moment.
[4] showed that within Cutibacterium acnes (then Propionibacterium acnes), distinct ribotypes (RT4, RT5, RT6) exist and differ in their association with acne. Function is read at strain resolution, not species resolution. Chapter 2 and Chapter 4 will carry this thread forward in detail.
[14] from the NIH/Segre lab demonstrated that an individual's skin metagenome is remarkably stable over 1–2 years. This finding placed the first quantitative constraint on the marketing language of "rebalancing your microbiome" — if a person's community is intrinsically stable, the claim that a single product can reset it within seven days carries a very high burden of evidence.
1.5 Setting the stage for AI — pre-AlphaFold ML for biology
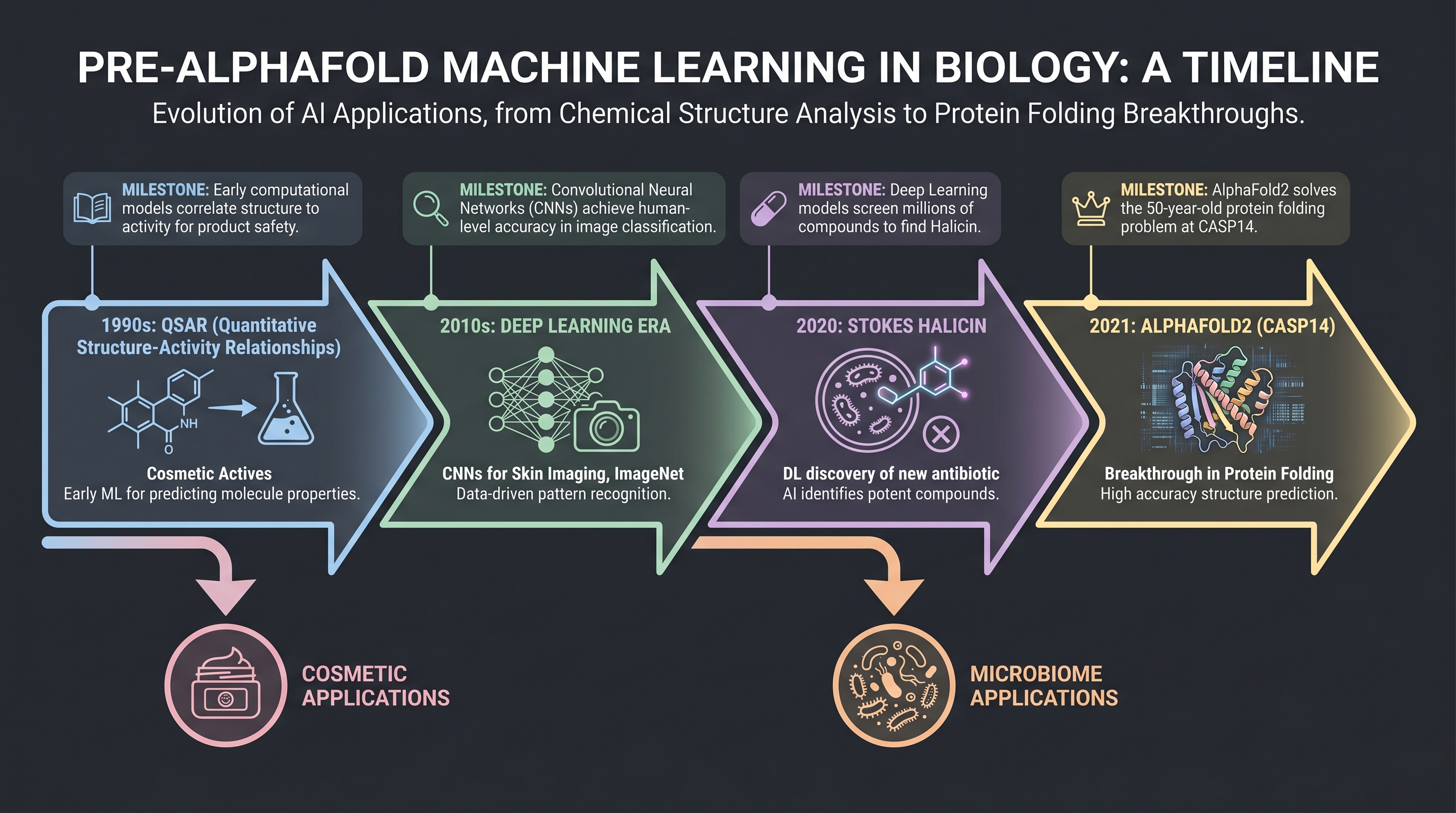
One central claim of this book is that AlphaFold 2 (2021) was a genuine inflection, but it did not arrive from nowhere (see Gap 9 in the gap map). Through the whole life of first-generation fermentation cosmetics, AI for biology was quietly accumulating elsewhere.
In the 1990s, QSAR (Quantitative Structure–Activity Relationship) and classical docking (e.g., AutoDock) became standard tools of drug design. In the 2000s, cheminformatics joined machine learning, and standard compound libraries — ChEMBL, ZINC — came into use. In the late 2010s, graph neural networks (GNNs) entered molecular structure prediction, and the apex of that line is [16] in Cell — the MIT Collins lab used a message-passing neural network (MPNN) to screen 110,000 molecules and discover halicin, a structurally novel antibiotic. This paper is often summarized as "the first AI-discovered antibiotic," but the substance is closer to: by 2020, AI could already identify new active molecules from within synthetic chemical libraries.
The reason this line of work did not flow into microbiome cosmetics at the time was data scarcity. The first systematic skin-microbe metabolite catalog had to wait for [13] (iHSMGC — integrated Human Skin Microbial Gene Catalog, 10.9M genes from a Chinese cohort). And the first tool that could predict the protein-metabolite interaction surface at industrial cost was [9], AlphaFold 2. That both appeared in the same year is not a coincidence — 2021 is the "year zero" that this book will return to repeatedly (Chapter 5 takes it up in full).
1.6 The paradigm shift — three axes moved at the same time
The transition from first-generation (fermentation) to second-generation (data-driven) cosmetics, drawn honestly as a research-paradigm shift rather than as a marketing slogan, runs along three axes.
Axis 1 — Single strain → ecosystem community. First-generation hypotheses were of the form "this strain produces this effect." Second-generation hypotheses are of the form "this ecological composition at this site produces this effect." [2] and [5] are the first measurements that made the transition possible.
Axis 2 — In vitro extract → ecosystem signaling. First-generation tests applied a ferment-filtrate extract to a single keratinocyte line. Second-generation tests ask how peptides secreted by a Cutibacterium acnes strain interact with the phenol-soluble modulins of Staphylococcus epidermidis to modulate AHR and NF-κB signaling in the skin's immune network — multi-omics, multi-species.
Axis 3 — Marketing claim → data-driven hypothesis. The "one hundred efficacies" language of the first generation was not falsifiable. A second-generation hypothesis is explicit — "applying this strain raises Cutibacterium RT6 relative abundance at the cheek by X% at week 4 baseline-corrected, and this change is r=Y correlated with transepidermal water loss (TEWL)." That explicitness is the surface on which AI can operate; AI cannot do anything with vague claims, only with measurable endpoints.
The three axes are coupled. The cost-curve collapse let researchers measure communities. Community measurement let researchers model ecosystem signaling. Modeling let measurable hypotheses become meaningful. The boundary between the first and second generations is not a single event but a five-year window (2009–2014) when three curves crossed their threshold together.
1.7 So is first-generation fermentation obsolete? — the book's first answer
The strongest question Brief A asks — "is fundamental research now obsolete?" — gets the same answer throughout the book: no, it is complementary. The full argument is in Chapter 3 and Chapter 9, but the skeleton has to be laid here.
What first-generation fermentation cosmetics did that remains valid has three parts. First, strain isolation, culture, and identification produced the wet-lab asset that is now AI's ground truth; AI predicts, culture validates. Prediction without validation is a demo, not a science. Second, antimicrobial and anti-inflammatory standard assays remain the foundation of endpoint definition; when an AI model calls a molecule "anti-inflammatory candidate," that label is ultimately defined by NF-κB, MMP, and IL-6 in vitro assays standardized since the 1980s. Third, clinical efficacy endpoints themselves (filaggrin expression, TEWL, collagen-synthesis rate, erythema index) are assets defined by the wet-lab era.
What first-generation fermentation cosmetics did that can no longer hold also has three parts. First, the "single fermentate vs vehicle" frame cannot capture strain resolution, site specificity, or temporal dynamics. Second, "one hundred efficacies" marketing-grade claims are becoming structurally indefensible under the FTC and EU CTR scrutiny of 2024–2026. Third, the industry pattern of operating for three decades without molecular mechanism — when AI now offers a feasible path to fill that mechanism — is no longer justifiable.
The reason this book ends in Chapter 12 with a research blueprint is precisely this: preserving the valuable assets of the first generation (strain libraries, assays, endpoints) while combining them with the data, AI, and synthetic biology of the second generation is the differentiation axis of the next decade of cosmetic R&D.
1.8 The Korean case — when industry catches up to mechanism
Korea is a special case of this paradigm shift. The country simultaneously holds dominant global market share in first-generation fermentation cosmetics (Sulwhasoo, The History of Whoo, SUM37) and a non-trivial share of second-generation data-driven research. [10], a Communications Biology paper from the COSMAX–Dankook–GIST collaboration, showed that the Lactobacillus rhamnosus IDCC 3201 strain secretes spermidine and that this spermidine drives reconstruction of collagen and elastin in human dermis. [11] published an EPI-7 clinical study in the International Journal of Molecular Sciences — postbiotics derived from the Epidermidibacterium keratini EPI-7 strain improved wrinkle and elasticity endpoints in a 12-week trial.
These two papers are perhaps the clearest examples of Korean fermentation cosmetics migrating from first-generation "appears effective" language to second-generation "this strain's this molecule moves this endpoint by this much" language. Chapter 10 takes up the Korean publication asymmetry directly — COSMAX is publication-visible while Amorepacific and LG H&H are not, even though both Big-2 firms are commercially leading.
1.9 One-page summary for the non-expert reader
If this chapter compresses to one page, the message is the following.
First-generation fermentation cosmetics (1980 — early 2010s) commercialized empirical efficacy in yeast and lactic-acid bacteria ferment filtrates, while the molecular mechanism of that efficacy remained largely unknown for thirty years. This was the era when marketing language ran three decades ahead of scientific language.
Between 2005 and 2014, sequencing cost dropped roughly a thousand-fold, and for the first time it became feasible to read the entire microbial ecosystem of a person's skin. Between 2009 and 2012, foundational papers — [2], [5], [3], [8] — followed in rapid succession, and skin microbiome research became "a field with instruments." This was microbiology's microscope moment.
In 2021, [9] (AlphaFold 2) and adjacent protein-structure-prediction tools made microbial-metabolite-to-skin-protein interaction prediction industrially feasible. This was the inflection at which AI could begin to compress the thirty-year mechanism gap of fermentation cosmetics. The remaining eleven chapters of this book are the anatomy of that inflection.
Fermentation is not finished. The new fermentation of the microbiome era is: data picks the strain, AI predicts the molecule, synthetic biology produces it, and the clinic validates it. Same word, a completely different industry.
1.10 Open Questions — three questions this book will return to
This chapter explicitly leaves three questions open for later chapters.
Open Question 1 — Are ferment-filtrate ingredients now obsolete? Galactomyces, Bifida, and Saccharomyces ferment filtrates are still on the shelves, and some ([17], [7]) have had their mechanisms retroactively explained. But many non-Galactomyces ferments remain in mechanism-deficit territory. The book's answer arrives in Chapter 5 (post-AlphaFold) and Chapter 7 (synthetic biology) — the future is less about ferment filtrates as such and more about identifying, purifying, and re-synthesizing the individual active molecules they contained.
Open Question 2 — Are wet-lab basics (strain isolation, culture, antimicrobial assays) outdated? The one-line answer is "no — they are more important now, as AI's ground truth." The full argument is in Chapter 3 (NGS, metagenomics, culturomics) and Chapter 9 (clinical efficacy validation).
Open Question 3 — How wide was the gap between first-generation marketing language and the underlying clinical data — and is the gap narrowing in the second generation? The answer runs through Chapter 8 (AI formulation), Chapter 10 (global firm case studies), and Chapter 12 (research blueprint) — without measurable endpoints and a reproducibility infrastructure, the second generation risks replaying the language asymmetry of the first.
These three questions recur to the final chapter. Without an honest assessment of the first generation's legacy, the second generation's promise will be overestimated. Without an honest assessment of the second generation's tools, the first generation's assets will be unjustly discarded. This chapter is the coordinate frame for both traps.
References
- Atallah, C., El Abiad, A., El Abiad, M., Nakad, M., & Assaf, J. C. (2025). Bioengineered Skin Microbiome: The Next Frontier in Personalized Cosmetics. Cosmetics, 12(5):205. doi:10.3390/cosmetics12050205.
- Costello, E. K., Lauber, C. L., Hamady, M., Fierer, N., Gordon, J. I., & Knight, R. (2009). Bacterial Community Variation in Human Body Habitats Across Space and Time. Science, 326(5960):1694–1697. doi:10.1126/science.1177486.
- Findley, K., Oh, J., Yang, J., Conlan, S., Deming, C., Meyer, J. A., Schoenfeld, D., Nomicos, E., Park, M., NISC Comparative Sequencing Program, Kong, H. H., & Segre, J. A. (2013). Topographic diversity of fungal and bacterial communities in human skin. Nature, 498(7454):367–370. doi:10.1038/nature12171.
- Fitz-Gibbon, S., Tomida, S., Chiu, B.-H., Nguyen, L., Du, C., Liu, M., Elashoff, D., Erfe, M. C., Loncaric, A., Kim, J., Modlin, R. L., Miller, J. F., Sodergren, E., Craft, N., Weinstock, G. M., & Li, H. (2013). Propionibacterium acnes strain populations in the human skin microbiome associated with acne35970-1/fulltext). Journal of Investigative Dermatology, 133(9):2152–2160. doi:10.1038/jid.2013.21.
- Grice, E. A., Kong, H. H., Conlan, S., Deming, C. B., Davis, J., Young, A. C., NISC Comparative Sequencing Program, Bouffard, G. G., Blakesley, R. W., Murray, P. R., Green, E. D., Turner, M. L., & Segre, J. A. (2009). Topographical and Temporal Diversity of the Human Skin Microbiome. Science, 324(5931):1190–1192. doi:10.1126/science.1171700.
- Grice, E. A., & Segre, J. A. (2011). The skin microbiome. Nature Reviews Microbiology, 9(4):244–253. doi:10.1038/nrmicro2537.
- Hashimoto-Hachiya, A., Furue, M., & Tsuji, G. (2022). Galactomyces Ferment Filtrate Potentiates an Anti-Inflammaging System in Keratinocytes. Journal of Clinical Medicine, 11(21):6338. doi:10.3390/jcm11216338.
- Human Microbiome Project Consortium (2012). Structure, function and diversity of the healthy human microbiome. Nature, 486(7402):207–214. doi:10.1038/nature11234.
- Jumper, J., Evans, R., Pritzel, A., Green, T., Figurnov, M., Ronneberger, O., Tunyasuvunakool, K., Bates, R., Žídek, A., Potapenko, A., et al. (2021). Highly accurate protein structure prediction with AlphaFold. Nature, 596(7873):583–589. doi:10.1038/s41586-021-03819-2.
- Kim, G., Kim, M., Kim, M., Park, C., Yoon, Y., Lim, D.-H., Yeo, H., Kang, S., Lee, Y.-G., Beak, N.-I., Lee, J., Kim, S., Kwon, J. Y., Choi, W. W., Lee, C., Yoon, K. W., Park, H., & Lee, D.-G. (2021). Spermidine-induced recovery of human dermal structure and barrier function by skin microbiome. Communications Biology, 4:231. doi:10.1038/s42003-020-01619-4.
- Kim, J., Lee, Y. I., Mun, S., Jeong, J., Lee, D.-G., Kim, M., Jo, H., Lee, S., Han, K., & Lee, J. H. (2023). Efficacy and Safety of Epidermidibacterium Keratini EPI-7 Derived Postbiotics in Skin Aging: A Prospective Clinical Study. International Journal of Molecular Sciences, 24(5):4634. doi:10.3390/ijms24054634.
- Kobyliak, N., Conte, C., Cammarota, G., Haley, A. P., Styriak, I., Gaspar, L., Fusek, J., Rodrigo, L., & Kruzliak, P. (2016). Probiotics in prevention and treatment of obesity: a critical view. Nutrition & Metabolism, 13:14. doi:10.1186/s12986-016-0067-0.
- Li, Z., Xia, J., Jiang, L., Tan, Y., An, Y., Zhu, X., Ruan, J., Chen, Z., Zhen, H., Ma, Y., Jie, Z., Xiao, L., Yang, H., Wang, J., Kristiansen, K., Xu, X., Jin, L., Nie, C., Krutmann, J., Liu, X., & Wang, J. (2021). Characterization of the human skin resistome and identification of two microbiota cutotypes. Microbiome, 9:47. doi:10.1186/s40168-020-00995-7.
- Oh, J., Byrd, A. L., Deming, C., Conlan, S., NISC Comparative Sequencing Program, Kong, H. H., & Segre, J. A. (2014). Biogeography and individuality shape function in the human skin metagenome. Nature, 514(7520):59–64. doi:10.1038/nature13786.
- Oh, J., Byrd, A. L., Park, M., NISC Comparative Sequencing Program, Kong, H. H., & Segre, J. A. (2016). Temporal Stability of the Human Skin Microbiome30399-3). Cell, 165(4):854–866. doi:10.1016/j.cell.2016.04.008.
- Stokes, J. M., Yang, K., Swanson, K., Jin, W., Cubillos-Ruiz, A., Donghia, N. M., MacNair, C. R., French, S., Carfrae, L. A., Bloom-Ackermann, Z., Tran, V. M., Chiappino-Pepe, A., Badran, A. H., Andrews, I. W., Chory, E. J., Church, G. M., Brown, E. D., Jaakkola, T. S., Barzilay, R., & Collins, J. J. (2020). A Deep Learning Approach to Antibiotic Discovery30102-1). Cell, 180(4):688–702.e13. doi:10.1016/j.cell.2020.01.021.
- Takei, K., Mitoma, C., Hashimoto-Hachiya, A., Tsuji, G., Takahara, M., & Furue, M. (2015). Galactomyces fermentation filtrate prevents T helper 2-mediated reduction of filaggrin in an aryl hydrocarbon receptor-dependent manner. Clinical and Experimental Dermatology, 40(7):786–793. doi:10.1111/ced.12635.