Chapter 4: AI-Driven Strain and Metabolite Screening
Why this chapter
If Part I showed what changed, Part II must answer why AI was the inflection point. The first answer lives in screening. The candidate space cosmetic-microbiome R&D now has to triage long ago surpassed what hands can validate — a single skin site holds hundreds of species and thousands of strains, each strain carries dozens to hundreds of biosynthetic gene clusters (BGCs), and each BGC potentially encodes a new metabolite. Every cosmetic endpoint (antimicrobial, anti-inflammatory, antioxidant, anti-aging) requires wet-lab assays costing tens to hundreds of dollars per candidate. Testing every candidate is arithmetically impossible.
Machine learning has a simple role: prioritize candidates before they reach the bench. Pharma has been doing this for thirty years — from Tanimoto-similarity hit triage to halicin's deep-learning discovery [25], to GENTRL-designed DDR1 inhibitors [29], to rentosertib in Phase 2a [22]. Cosmetic-microbiome is adopting the same tools five to ten years behind, but a naive port does not work — the shape of the domain data differs. This chapter maps that gap.
Three quantitative anchors for this chapter 1. ML pipeline shape per endpoint: molecular fingerprint → classical ML → graph/transformer model. Inflection points: [25]'s MPNN, [3]'s SHAP-based XAI. 2. Data-source asymmetry: metabolite side is public-functional (NPAtlas, MIBiG, GNPS); strain × clinical-endpoint side is effectively private (Unilever 30K samples, COSMAX FACE-LINK 950 subjects). This asymmetry caps model generalization. 3. Five industry cases: Insilico (pharma), Cradle and Profluent (protein-design SaaS), COSMAX × Dankook FACE-LINK (Korean cosmetics), L'Oréal × Lactobio (data-via-M&A), Unilever (internal XAI pipeline). The cosmetic industry has yet to publish a peer-reviewed clinical trial of an AI-designed active ingredient — Gap 1.
4.1 The screening bottleneck — pre-AI numbers
In the fermentation era (Chapter 1), discovery of cosmetic actives was a mixture of experience, accident, and reverse engineering. SK-II's Pitera began with the hands of sake-brewery workers; the molecular mechanism of Galactomyces ferment filtrate as an AHR agonist was only resolved in 2015 — three decades after the product reached market [26]. With the metagenomics era (Chapter 3), the candidate space exploded. The single iHSMGC skin metagenomic catalog alone registered 10.94 million genes, of which roughly 45% were invisible to HMP [16].
The most intuitive scale unit is the BGC count. [2] highlights that antiSMASH-identified BGCs across the skin, oral, and gut microbiome reach hundreds per individual on average, with even more uncharacterized RiPP (ribosomally synthesized post-translationally modified peptide), NRPS (non-ribosomal peptide synthetase), and PKS (polyketide synthase) clusters at the species level. Following one cluster from sequence to isolated product, structural determination, and assay typically consumes 6–24 months of a PhD-grade researcher's time. Tracking every BGC at parity is staffing-arithmetic impossible.
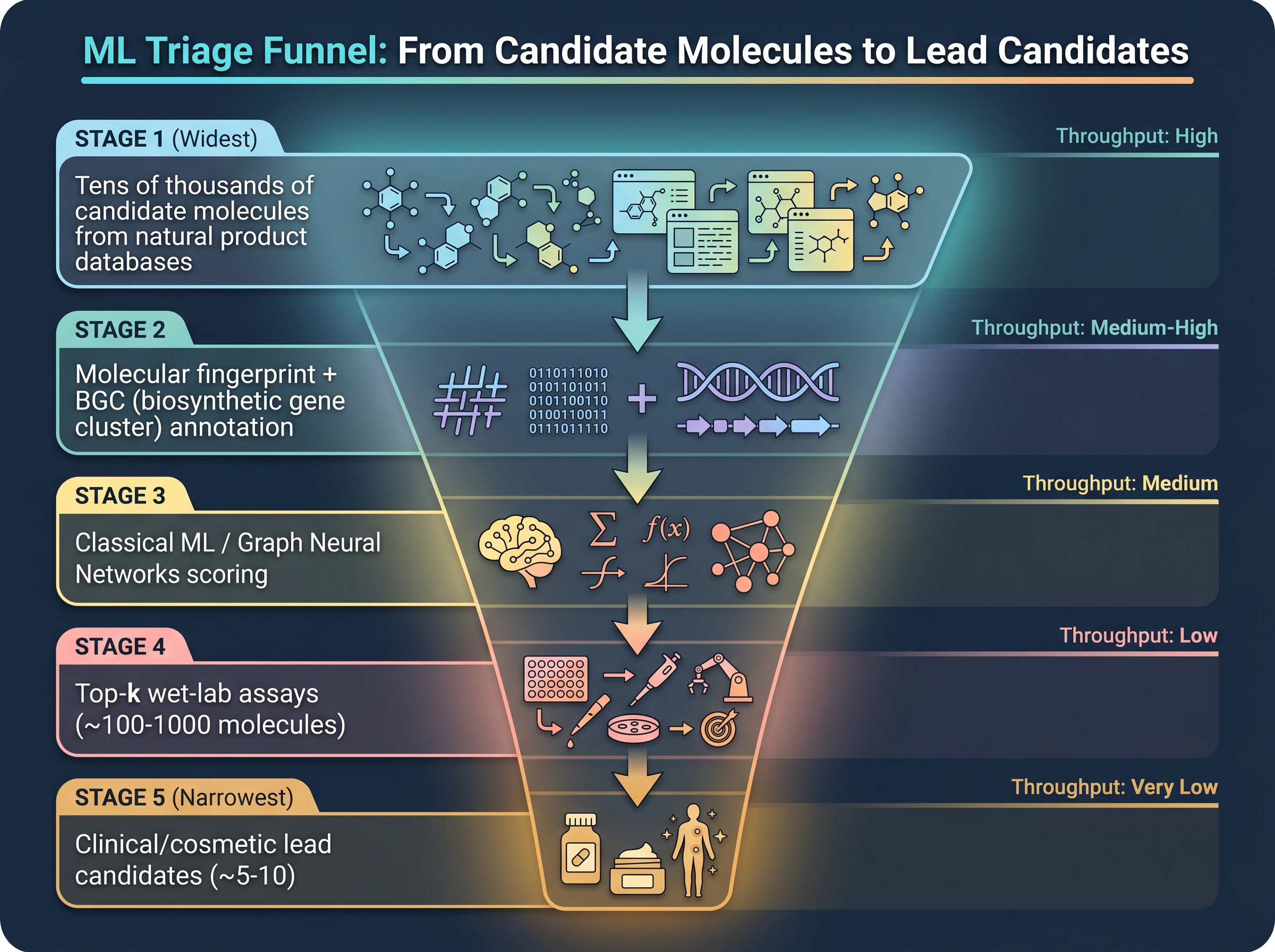
But random downsampling crushes the hit rate. [25]'s halicin study leveraged this asymmetry precisely — a message-passing neural network (MPNN) was trained on a 2,335-molecule primary library, then used to score the 6,111-molecule Drug Repurposing Hub (a curated set of FDA-approved and clinical-stage drugs). The top 99 model-ranked candidates were tested empirically against E. coli, and 51 (51.5%) were confirmed active — a far higher hit rate than random library screens. After downstream filtering for structural diversity, toxicity, and similarity to known antibiotics, roughly 1–3 leads remained, with halicin the most prominent. That number is for antibiotics — the best-defined endpoint. Cosmetic endpoints — "anti-aging efficacy," "sensitive-skin soothing" — are harder to reduce to a single in vitro assay, so baseline hit rates are likely lower. Exact figures are locked inside industry [8].
The promise of ML triage is to push a noisy library's sub-1% hit rate to 30–50% within the top-ranked candidate pool — the halicin study's 51.5% (51 active among the top 99) is tens of times the random-screen baseline, and a yield at which wet-lab throughput can meaningfully absorb the ML output. No cosmetic study with comparable external validation has been published — Gap 1, discussed below.
4.2 An endpoint catalog — same tools, different proxies
The largest reason cosmetic ML differs from pharma ML is that endpoint definitions are weaker. Antibiotic activity reduces to MIC (minimum inhibitory concentration), a single in vitro reading. "Anti-aging" does not. Some endpoints admit simplification; others do not — and that division determines what data scale a model can rely on.
Antimicrobial — the best-defined endpoint. MIC and MBC (minimum bactericidal concentration) against E. coli, S. aureus, C. acnes serve as common currency. [25]'s MPNN ports directly. [15] reports that the commensal Staphylococcus epidermidis produces a polyene peptide called epifadin that clears nasal S. aureus — a clean molecular template of the "microbe versus microbe" concept already commercialized in cosmetics. [24] extends this stream to the cosmetic side, becoming the first AI tool predicting skin-microbiome-mediated metabolism of biotics and xenobiotics.
Anti-inflammatory — proxy assays diverge. NF-κB and AP-1 reporter cell assays, IL-6/IL-8/TNF-α suppression, AHR and NRF2 activation. [9] added a second molecular pillar by demonstrating NRF2 activation for Galactomyces ferment filtrate. Model inputs are usually molecular fingerprints or peptide sequences, outputs are quantitative scores from multiple reporter assays, and the mapping differs per endpoint — which is why multi-task learning becomes the natural architecture.
Antioxidant — DPPH, ABTS, ORAC, and cell-based ROS readouts. QSAR models predicting ORAC scores from molecular fingerprints have been stable since the 1990s; deep learning has only nudged the accuracy. In cosmetic ML, this is the most commoditized endpoint.
Anti-aging — fundamentally a composite endpoint. Collagen synthesis (procollagen ELISA), MMP-1 suppression, fibroblast senescence markers (SA-β-gal, p16), with the in vivo aggregate of dermal density and wrinkle-depth measurements. No single in silico endpoint suffices, so multi-omics integration (transcriptomics + metabolomics + clinical imaging) becomes the dominant training signal. [28]'s review of skin microbiome and aging makes the causal-unsolvedness explicit — correlations are plentiful, mechanistic chains only partially worked out.
Table 4.1 — ML input, output, and generalization ceiling per endpoint | endpoint | model input | target output | public data scale | generalization ceiling | |---|---|---|---|---| | Antimicrobial (MIC) | molecular graph, peptide sequence | MIC ≤ threshold (binary) | ChEMBL, PubChem, MIBiG | strong — external validation feasible | | Anti-inflammatory (NF-κB) | molecular fingerprint | reporter suppression (regression) | medium, partly public | moderate — assay variance | | Antioxidant (ORAC) | molecular fingerprint | ORAC unit (regression) | medium, mostly public | strong — most commoditized | | Anti-aging (collagen) | molecular + multi-omics | procollagen + MMP-1 + clinical imaging | effectively private | low — internal data dependence |
This table is where Gap 2 (no skin-microbiome foundation model) and Gap 12 (no open benchmarks) originate — for the cosmetic-core endpoints (anti-aging, anti-inflammatory), data is private, models are retrained inside firms, and external comparability disappears [20].
4.3 Data sources — metabolite-public, strain-private
The data terrain of cosmetic-microbiome ML splits cleanly in two.
Metabolite (public asset)
- NPAtlas — a public catalog of microbial and marine natural-product molecules. As of 2025 it indexes tens of thousands of molecules with SMILES, MOL, and structural classification — the most common starting point for molecular-fingerprint training in cosmetic ML.
- MIBiG (Minimum Information about a Biosynthetic Gene cluster) — curated links between BGCs and their products. Starting from antiSMASH annotation and tracing all the way to product structures, MIBiG provides the ground-truth set [2] highlights as essential infrastructure for cosmetic-grade RiPP/NRPS search.
- GNPS (Global Natural Products Social) — public MS/MS spectrum repository. The standard library cosmetic firms use when mapping their internal metabolomics to NPAtlas/MIBiG.
- FAST-NPS — a high-throughput automated genome-mining pipeline published in Cell Systems [7]. From BGC to product to MS-guided purification in a single track, demonstrated on Streptomyces — porting to cosmetic-relevant taxa (S. epidermidis, C. acnes, Cutibacterium) is the obvious next step.
This metabolite infrastructure is, by academic-industry collaboration, comparatively open, and molecular-fingerprint ML (MPNN, Graph Transformer) can train on it directly.
Strain × clinical endpoint (private asset)
- HMP / iHSMGC / MGnify — taxonomic information and gene catalogs are public, but metadata mapping to cosmetic endpoints (skin imaging, clinical scoring) is usually absent.
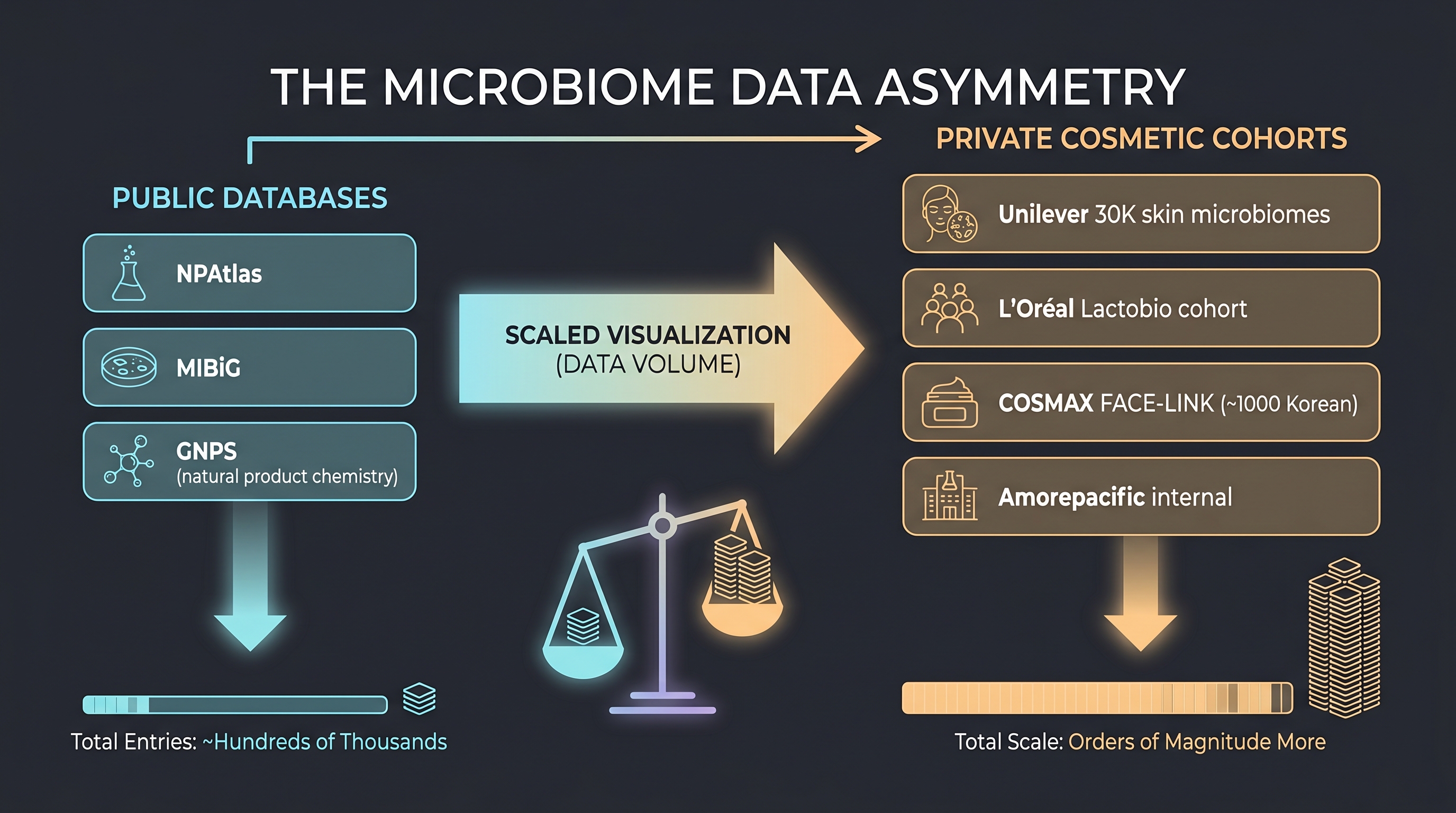
- Unilever internal data — [27] discloses a 30,000-sample internal cohort with 5 billion data points. No external access.
- L'Oréal internal data — Modjoul and Modiface device data plus the ~10,000-isolate culture collection absorbed via the Lactobio acquisition [17].
- COSMAX × Dankook FACE-LINK — a 950-subject Korean cohort with skin-microbiome ↔ biophysical mapping [18]. The flagship example of a Korean-style internal cohort, treated in detail in Chapter 10. (Note: "FACE-LINK" is COSMAX's internal platform/product brand name; the [18] paper itself does not use this label — this book pairs the two as the academic and commercial faces of the same cohort, following COSMAX's public communications.)
- EPI-7 clinical — [13] reports an 8-week RCT of postbiotics derived from Epidermidibacterium keratini EPI-7 — a rare example of a strain × clinical-endpoint mapping that reached peer review.
The asymmetry is explicit — academic ML progress is possible on the metabolite side, but "which strain improves which endpoint in which person" cannot be learned without company-internal data. [20]'s ML-for-microbiome guidelines emphasize batch effects, compositionality, kit bias — and these problems double over private cohorts. Among 200+ AI-microbiome papers consolidated by [14], externally validated models mapped to skin endpoints are rare.
4.4 Classical ML baselines — thirty-year tools that did not disappear
Hidden in the shadow of AI hype, the working baseline of cosmetic-microbiome ML is still random forests and gradient boosting. The reason is simple — per-endpoint training data sits at hundreds to thousands of samples, and at that scale deep learning does not trivially win. [3]'s IBM × Unilever collaboration is the case study. To predict phenotype (skin condition) differences from 16S data of a ~50-subject cohort, they used gradient boosting plus SHAP (Shapley value–based explainability). They did not bolt on a CNN or Transformer not from theoretical elegance but from data scale — fitting a deep model on a 50-subject cohort guarantees overfit.
Tanimoto similarity-based hit prioritization is similarly the chemoinformatics standard. Representing molecular fingerprints (Morgan, MACCS, RDKit) as bit vectors and filtering candidates by Tanimoto distance < 0.3 from known actives (e.g., a known anti-aging peptide) is — unglamorous, but — the fastest narrowing of the wet-lab queue.
Another reason classical ML dominates cosmetic R&D: interpretability. Backing a marketing claim with "deep learning predicted so" rarely passes regulatory or legal review. Variable-importance outputs of SHAP/LIME/feature importance translate to R&D-meeting decisions, and cosmetic industry is, on this point, more conservative than pharma [6].
4.5 Sequence-to-function neural models — the bridge before and after AlphaFold
Starting endpoint prediction from molecular fingerprints works when the molecule is already isolated. The true ambition of microbiome ML is to go from metagenomic reads straight to efficacy candidate molecules. The core algorithms for that flow stabilized in the early-to-mid 2020s.
DeepBGC, antiSMASH + ML — models that detect BGC regions in metagenomic reads or assembled contigs. antiSMASH started rule-based (hidden Markov models); DeepBGC raised accuracy with word2vec embeddings plus bidirectional LSTM in 2019. The output is "this contig contains a BGC, the product is likely of class X (RiPP/NRPS/PKS/terpene…)" — directly usable in cosmetic R&D to rank a strain collection by BGC density. [7] is the industrialization of this flow — Streptomyces genome → BGC → product candidates in one pipeline.
RFM (Retrosynthesis & Function Mining) — metagenome → peptide → efficacy. The stream is most mature for antimicrobial peptides (AMPs). Embed peptide sequences with a protein language model like ESM-2, fine-tune an antimicrobial-activity classifier. For cosmetics, finding more lugdunin-class or epifadin-class peptides [15] across similar commensal strains is the near-term industrial task.
Graph neural networks for community function — the stream of predicting function at the community level rather than the single strain. Inter-species interactions are encoded as graphs, and message passing predicts community-level metabolite production. Discussed further in Chapter 6 in the digital-twin context, this is — as [14] and [5] both stress — the most active academic stream right now, but clinical-endpoint-mapped validations remain rare.
A common limit: ground truth for validation remains wet-lab. Even if ML predicts a BGC class, the exact product structure must be determined by NMR/MS, and peptide antimicrobial activity must be confirmed on plate assays. This dovetails exactly with Chapter 3's message — fundamental research is not obsolete; AI accelerates, but validation is still hands-on.
4.6 Industry cases — five companies, five messages
4.6.1 Insilico Medicine — the clinical readout cosmetics has not had
Insilico Medicine is not a cosmetic firm, but the only company that has demonstrated a peer-reviewed clinical readout of an AI-designed active ingredient. In 2019, GENTRL [29] used a reinforcement-learning–based generative model to take DDR1 kinase inhibitor candidates from design to in vitro validation in ~46 days — a one-shot paper, but one that announced a new slope on the discovery-cost curve. The same group's PandaOmics platform [21] later integrated target identification through drug-candidate selection in a unified workflow.
The decisive readout came in 2024 with rentosertib (ISM001-055). [22], in Nature Biotechnology, reported the end-to-end generative-AI discovery — target identification (PandaOmics) → molecular design (GENTRL/Chemistry42) → Phase 1, in 30 months. Following this, [12] in Nature Medicine reported Phase 2a results — a TNIK inhibitor for idiopathic pulmonary fibrosis with FVC at 12 weeks of +98.4 mL versus −20.3 mL placebo. This is one of the first peer-reviewed clinical readouts the AI-drug-discovery field has produced — a readout the cosmetic industry does not have.
The cosmetic implication: to run the same workflow (target → molecular design → clinical) for a cosmetic endpoint (anti-aging, soothing), endpoint definitions need to be tightened sufficiently, and the capital and time discipline to push to clinical readout must be present. Chapter 9 dissects the readout template, and Chapter 12 integrates it into the industry-wide blueprint as a decision variable.
4.6.2 Cradle Bio, Profluent — SaaSification of protein design
Among cosmetic actives, proteins and peptides are the candidates of the new era. Vegan collagen, bioactive peptides, enzyme-form actives — all reduce to protein design problems. Cradle Bio (Zurich/Delft) provides generative protein design as SaaS, closing a $73M Series B in November 2024 [4]; around the same time [19] publicly disclosed Cradle as a customer — the first proof that pharma majors do not build every in-house model and rely on external platforms.
Profluent drew more attention with [23]'s bioRxiv release of OpenCRISPR-1 — a ProGen2-based language model trained on the universe of CRISPR-Cas proteins, generating new genome editors and open-sourcing part of the catalog. Cradle is closed-SaaS, Profluent is open-plus-commercial — both business models are currently surviving the protein-design market.
The cosmetic implication: cosmetic firms do not have to build protein-design infrastructure in-house. Outsourcing vegan collagen or peptide actives to external SaaS directly compresses timelines, and industry velocity has reached the rhythm of cosmetic launch cycles (typically 18–24 months). Covered further in Chapter 7 alongside the synthetic-biology DBTL loop.
4.6.3 COSMAX × Dankook FACE-LINK — Korea's first published readout of an internal cohort
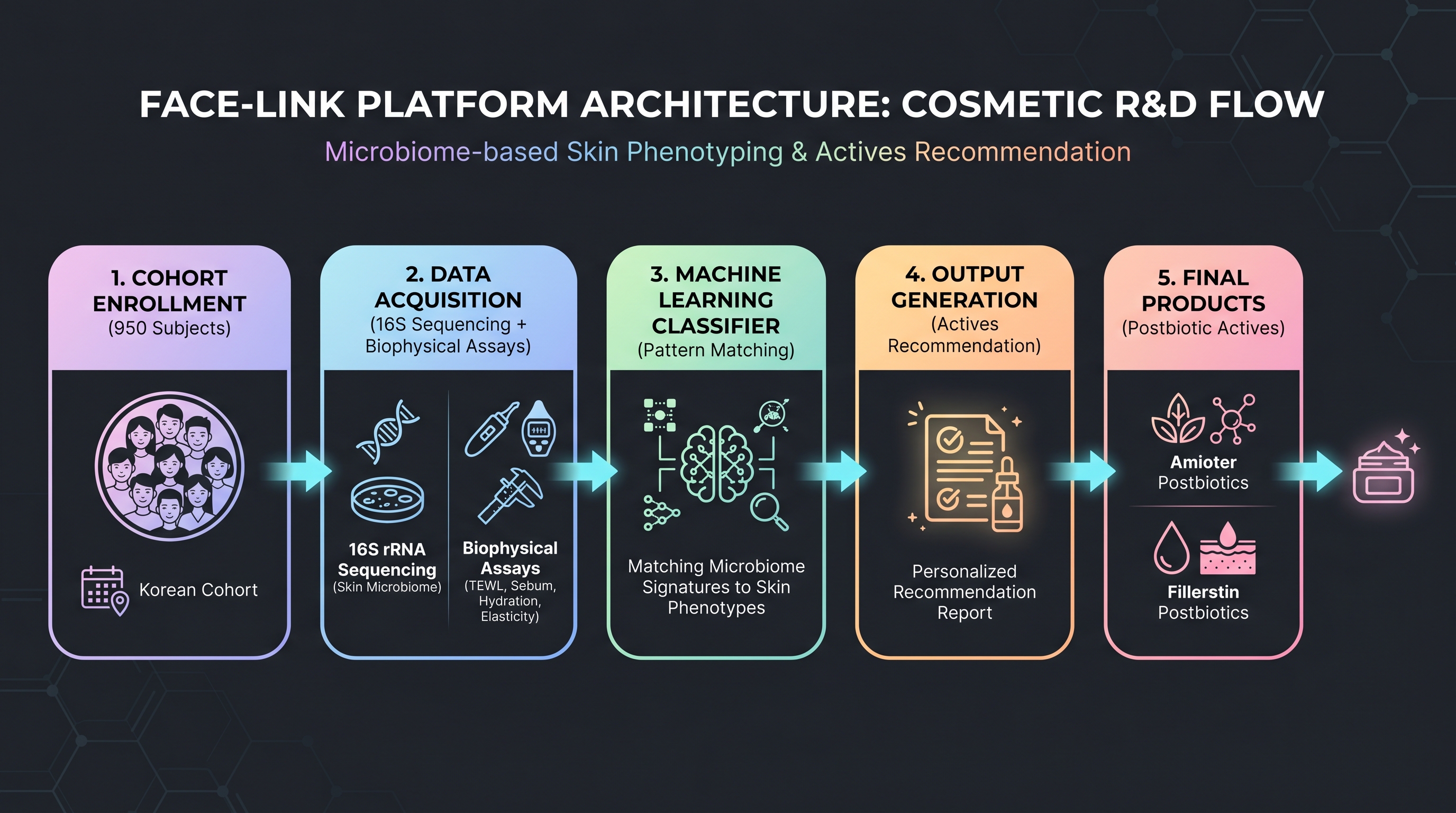
[18], in Frontiers in Cellular and Infection Microbiology, published an integrative analysis of a 950-subject Korean cohort mapping skin microbiome to biophysical measurements — FACE-LINK's first peer-reviewed readout. The 16S data is accompanied by skin hydration, TEWL (transepidermal water loss), and melanin/erythema readings matched per individual, and the paper validates an ML classifier that groups Korean skin types and aging brackets by microbiome signature.
What matters here is not data scale but data structure — most Western academic cohorts collect either 16S or metabolomics alone, while FACE-LINK matches microbiome and biophysical metrics within the same individual. Such matched cohorts are the foundation of supervised learning, so models can be trained directly against cosmetic endpoints.
Korea's structural asset: COSMAX is a B2B OEM, so publication is a commercial signal — different from Western cosmetic firms with no peer OEM. Amorepacific and LG H&H are B2C brand owners, with weaker incentives to publish (Gap 4). FACE-LINK is, as a result, almost the only externally visible internal cohort from Korea. Chapter 10 places this within the broader Korean industry matrix.
4.6.4 L'Oréal × Lactobio — buying the data
L'Oréal acquired the Danish microbiome firm Lactobio in December 2023 [17], closing in Q1 2024. Lactobio brought two assets: (1) a culture collection of ~10,000 isolated and characterized strains, (2) the corresponding efficacy data (mostly antimicrobial and anti-inflammatory in vitro). L'Oréal R&I is now combining this strain asset with its prior molecular and peptide AI pipelines — together with Modjoul/Modiface device data covered in Chapter 10 — to build multi-modal cosmetic-microbiome R&D infrastructure.
The strategic message: in cosmetic-microbiome, buying the data is faster than growing the AI. Model architectures get commoditized fast by academic AI, but strain collections and clinical-endpoint mappings do not. The Lactobio acquisition is a rational response to that asymmetry — and subsequent industry M&A has used it as the template.
4.6.5 Unilever — standardizing the internal XAI pipeline
[3] is a Scientific Reports paper co-authored by Unilever R&D and IBM Research, reporting gradient boosting plus SHAP to predict phenotype differences from skin microbiome composition. The paper is one of the first cases of methodologically responsible ML application from cosmetic industry — an n=1 academic publication, but interpretable as the standardization origin of Unilever's internal pipeline.
In 2025 [27] disclosed at SXSW and on their R&D page — 30,000 samples, 5 billion data points internal cohort, AI virtual cohorts (2,500 simulated subjects), formulation cycles compressed from 5–6 to 1–2, and claim development accelerated 75%. Operational KPIs are striking, but external validation is absent (Gap 15). [10]'s PRISMA review of 74 cosmetogenomics-AI studies explicitly notes that no firm, including L'Oréal and Unilever, has published a peer-reviewed clinical readout of an AI-designed cosmetic active.
The gap between Carrieri 2021's external visibility and Unilever 2025's internal KPIs is Gap 1 — AI is heavily used in cosmetic-industry discovery, triage, and formulation, but no readout reaching clinical depth has yet appeared in peer-reviewed channels.
4.7 The gap — Insilico's readout, cosmetics' absence (linking Chapters 9 and 12)
This chapter's single most important line: as of May 2026, no peer-reviewed clinical trial of an AI-designed cosmetic active ingredient exists. The toolkit is ready — AlphaFold 3 [1], ESM3 [11], generative molecular design, BGC mining are all commodity by 2024. The data, though private, is sufficient — Unilever 30K, COSMAX FACE-LINK 950, L'Oréal Lactobio 10K strains. What is missing is industry incentive.
[10] articulates two reasons. (1) Cosmetic actives do not require FDA approval at drug-grade evidentiary depth, so the commercial ROI of a peer-reviewed clinical readout is low — well-designed marketing claims are more efficient. (2) A successful AI-designed active immediately becomes patent IP, deleting the publication incentive. [2] puts it bluntly — "efficacy claims for bioengineered actives rely on company press releases, not peer-reviewed clinical trials."
The R&D-planning implication: the first firm to publish such a readout — even if the active is not an immediate marketing success — gains a category-defining asset. Among Korean firms (COSMAX, Amorepacific, LG H&H) or a smaller EU player (pre-acquisition Lactobio-style outfits), the first publishable readout looks likely. Chapter 9 details the clinical readout template, and Chapter 12 integrates it as a decision variable in the industry-wide blueprint.
4.8 Open Questions
- Endpoint proxy validity — How tightly do procollagen ELISA and MMP-1 expression in anti-aging readouts correlate causally with in vivo dermal density and wrinkle depth? External studies quantifying the clinical correlation of the proxies cosmetic ML trains on are rare.
- Data-scale ceiling — At what cohort size does a single-phenotype internal dataset plateau? Unilever 30K and COSMAX FACE-LINK 950 do not let outsiders estimate where the learning curve flattens (Gap 2). If it flattens before 100K, the industry's data-hoarding race would not be rational.
- Skin-type and ethnicity generalization — [10] flags "limited geographic diversity, darker phototypes underrepresented" as structural problems. Models trained on Korean (FACE-LINK), Chinese (iHSMGC), or Western (HMP) cohorts — how badly do they generalize across ethnicities? Directly affects global cosmetic launches.
- BGC-class bias — RiPP, NRPS, and PKS classes that current ML triage handles well are the classes for which training data is rich. Variants more common in cosmetic-core taxa (Cutibacterium, Malassezia) — including unusual terpene synthases — may be exactly where models are weakest, which is also where novel discovery lives.
- Validation-resource distribution — Even if ML successfully reduces 100 candidates to 5, taking those 5 to clinical readout is a different staffing-capital problem. If cosmetic R&D's wet-lab throughput cannot absorb the ML output — the likely industry scenario — where does the next bottleneck move?
References
- Abramson, J., Adler, J., Dunger, J. et al. (2024). Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500.
- Atallah, C., El Abiad, A., El Abiad, M. et al. (2025). Bioengineered Skin Microbiome: The Next Frontier in Personalized Cosmetics. Cosmetics 12(5):205.
- Carrieri, A. P., Haiminen, N., Maudsley-Barton, S. et al. (2021). Explainable AI reveals changes in skin microbiome composition linked to phenotypic differences. Scientific Reports 11:4565.
- Cradle Bio (2024). Cradle's $73M Series B for AI protein design. TechCrunch / SynBioBeta, Nov 2024. [Cradle, 2024]
- Deep learning microbiome review — Wang, T., Yang, L. et al. (2024). Deep learning in microbiome analysis: a comprehensive review of neural network models. Frontiers in Microbiology 15:1516667.
- Di Guardo, A., Trovato, F., Cantisani, C. et al. (2025). Artificial Intelligence in Cosmetic Formulation: Predictive Modeling for Safety, Tolerability, and Regulatory Perspectives. Cosmetics 12(4):157.
- FAST-NPS authors (2025). FAST-NPS — high-throughput automated genome mining for bioactive natural products00040-X). Cell Systems, March 2025. [FAST-NPS, 2025]
- Gueniche, A., Perin, O., Bouslimani, A. et al. (2022). Advances in Microbiome-Derived Solutions and Methodologies Are Founding a New Era in Skin Health and Care. Pathogens 11(2):121.
- Hashimoto, K., Yamamoto, T., Yagi, M. et al. (2022). NRF2 activation by Galactomyces ferment filtrate complements the AHR-axis mechanism in skin barrier homeostasis. Journal of Cosmetic Dermatology, 2022. [Hashimoto et al., 2022]
- Haykal, D., Flament, F., Amar, D. et al. (2025). Cosmetogenomics unveiled: a systematic review of AI, genomics, and the future of personalized skincare. Frontiers in Artificial Intelligence 8:1660356.
- Hayes, T., Rao, R., Akin, H. et al. (2025). Simulating 500 million years of evolution with a language model (ESM3). Science, 2025.
- Insilico Medicine clinical authors, Ren, F., Zhavoronkov, A. et al. (2025). A generative AI-discovered TNIK inhibitor for idiopathic pulmonary fibrosis: a randomized phase 2a trial. Nature Medicine, May 2025. [Insilico, 2025]
- Kim, J., Lee, Y. I., Mun, S. et al. (2023). Efficacy and Safety of Epidermidibacterium Keratini EPI-7 Derived Postbiotics in Skin Aging: A Prospective Clinical Study. International Journal of Molecular Sciences 24(5):4634.
- Wang, X.-W., Wang, T., Liu, Y.-Y. (2024). Artificial Intelligence for Microbiology and Microbiome Research. arXiv preprint 2411.01098. [Wang et al., 2024]
- Krismer, B., Peschel, A. et al. (2024). Commensal production of broad-spectrum antimicrobial peptide polyene eliminates nasal S. aureus (epifadin). Nature Microbiology, 2024.
- Li, Z., Xia, J., Jiang, L. et al. (2021). Characterization of the human skin resistome and identification of two microbiota cutotypes (iHSMGC catalog). Microbiome 9:47.
- L'Oréal R&I Press (2023). L'Oréal acquires Lactobio to strengthen microbiome research. L'Oréal press, Dec 2023; closed Q1 2024. [L'Oréal, 2023]
- Mun, S., Jo, H., Heo, Y. M. et al. (2025). Skin microbiome-biophysical association: a first integrative approach to classifying Korean skin types and aging groups (FACE-LINK). Frontiers in Cellular and Infection Microbiology 15:1561590.
- Novo Nordisk; Cradle (2024). Novo Nordisk × Cradle AI protein design partnership disclosure. Cradle Series B press, Nov 2024. [Novo Nordisk, 2024]
- Papoutsoglou, G., Tarazona, S., Lopes, M. B. et al. (2023). Machine learning approaches in microbiome research: challenges and best practices. Frontiers in Microbiology 14:1261889.
- Pun, F. W., Ozerov, I. V., Zhavoronkov, A. et al. (2024). PandaOmics: An AI-Driven Platform for Therapeutic Target and Biomarker Discovery. Journal of Chemical Information and Modeling, Feb 2024.
- Ren, F., Ding, X., Zheng, M. et al. (2024). A small-molecule TNIK inhibitor (ISM001-055 / rentosertib) discovered via end-to-end generative AI from target identification to Phase 1. Nature Biotechnology, Mar 2024.
- Ruffolo, J. A., Nayfach, S., Gallagher, J. et al. (2024). Design of highly functional genome editors by modeling the universe of CRISPR-Cas proteins (Profluent ProGen2 / OpenCRISPR-1). bioRxiv 2024.04.22.590591.
- Skin Bug authors — Lee, S. et al. (2021). SkinBug — AI prediction of skin microbiome-mediated metabolism of biotics and xenobiotics. iScience 24(1):102026. [SkinBug, 2021]
- Stokes, J. M., Yang, K., Swanson, K. et al. (2020). A Deep Learning Approach to Antibiotic Discovery30102-1). Cell 180(4):688–702.
- Takei, K., Mitoma, C., Hashimoto-Hachiya, A. et al. (2015). Galactomyces ferment filtrate as AHR agonist restoring filaggrin and skin barrier function. Journal of Dermatological Science, 2015. [Takei et al., 2015]
- Unilever Beauty & Wellbeing R&D (2025). How Unilever's pioneering skin microbiome research is shaping product innovation. Unilever news; SXSW 2025 + R&D page coverage. [Unilever, 2025]
- Wang, Y., Liu, R., Chen, S. et al. (2025). Skin microbiome and aging: a review of clinical and molecular evidence. Microbiological Research, 2025. [Wang et al., 2025]
- Zhavoronkov, A., Ivanenkov, Y. A., Aliper, A. et al. (2019). Deep learning enables rapid identification of potent DDR1 kinase inhibitors. Nature Biotechnology 37, 1038–1040.