Chapter 6: Microbiome–Skin Interaction Modeling and Digital Twins
Why this chapter
If (Chapter 5) was about predicting one protein × one metabolite binding, this chapter is a different arithmetic — a many-body problem: dozens of microbes × dozens of host cell types × time. The questions cosmetic R&D actually faces almost always live on this side. "If we apply this strain, how does the C. acnes–S. epidermidis ratio shift over a month?" "Which node along the skin-barrier recovery pathway is the L. ferment filtrate acting on?" "Why does the same active peptide behave differently on oily vs dry skin?" None of these reduce to a docking score between two molecules. They demand modeling of community-level dynamics.
This chapter maps where that modeling stands in 2026. Graph neural networks (GNNs), mechanistic dynamical models (Lotka–Volterra, agent-based, ODE), multi-omics integration, and the industry concept that goes by the name "digital twin." One message has to be stated up front and held throughout: as of May 2026, no skin-microbiome foundation model exists at ESM3 scale. This is the gap the critical-analyst flagged as Gap 2 for the book, and it is the single most load-bearing finding of this chapter.
Three quantitative anchors for this chapter 1. Two streams of community modeling — data-driven GNNs (SIMBA-GNN, [24]) and mechanistic ODE / agent-based models (Lotka–Volterra extensions, in vitro microecology). They are complementary. 2. Data asymmetry — Unilever's internal 30,000-sample cohort and the COSMAX FACE-LINK 950-subject cohort ([19]) are large enough to technically train a foundation model. Neither has been opened externally. 3. The digital-twin definition gap — L'Oréal Skin Genius, Unilever virtual cohorts (2,500 simulated subjects), POND's Skin Institute analyzer ([23]) all make commercial twin claims. Methodologically validated twins (prediction + measurement + error quantification + external generalization) are not yet in the peer-reviewed channel ([7]).
6.1 The community problem — pairwise prediction is not enough
The protein–ligand docking models in (Chapter 5) carry one big assumption: two-body equilibrium. Fix the binding affinity between two molecules and you can estimate efficacy. The microbiome rarely obliges.
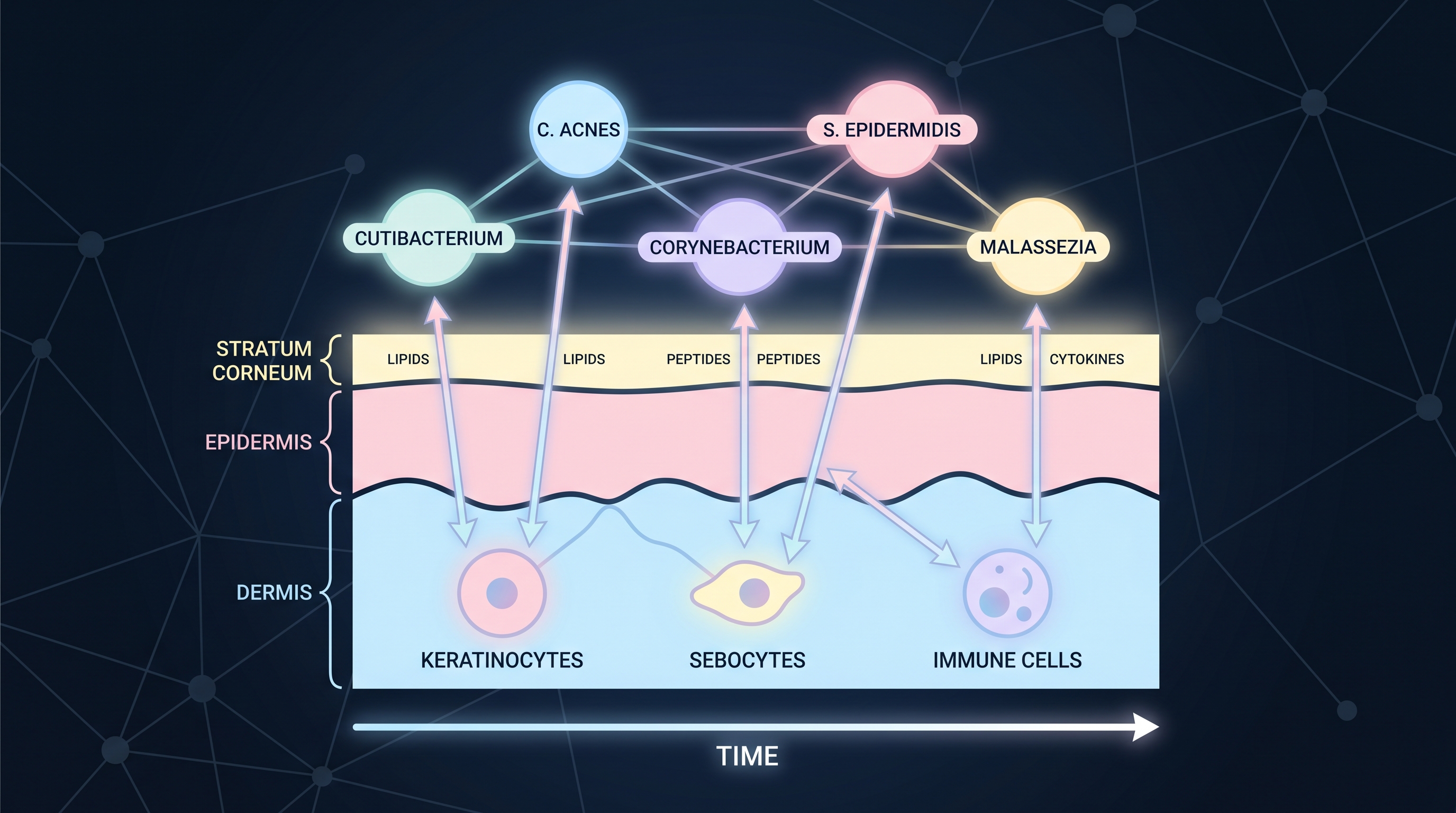
Even within a single skin site, Cutibacterium acnes is almost never acting alone — neighboring S. epidermidis may be releasing an epifadin-class polyene peptide [13], while nearby Malassezia hydrolyzes lipids to release free fatty acids, those free fatty acids feed C. acnes dysbiosis, and host keratinocytes respond through NF-κB and AHR ([3]). [11] showed that S. epidermidis, classed as commensal on healthy skin, in fact exacerbates inflammation on a damaged barrier — making clear that "strain → efficacy" is not a property of the strain's identity but of its context. Combined with the strain-level dynamics covered in (Chapter 2) ([16]), the picture is this: even within a species, strain matters, and even within a strain, what the bug does depends on the neighboring community and host state.
Reducing this many-body system to a sum of pairwise predictions fails for two reasons. First, nonlinear interactions — three microbes in proximity produce metabolic output that is not the sum of the three pairwise edges. Second, time dependence — C. acnes type IA1 → II transitions take weeks, while S. epidermidis stays roughly stable in the same window ([21]). A prediction calibrated on a single-time-point snapshot breaks at the next time point.
The starting point for community modeling is a network representation. Species (or strains) become nodes, interactions become edges, and edge weights are either learned from data (data-driven) or specified by differential equations (mechanistic). Lay host cells on top as additional nodes and you get a host–microbe interaction network; lay time on top and you get a dynamical system. This chapter walks down that stack.
6.2 Graph neural networks — learning edges from data
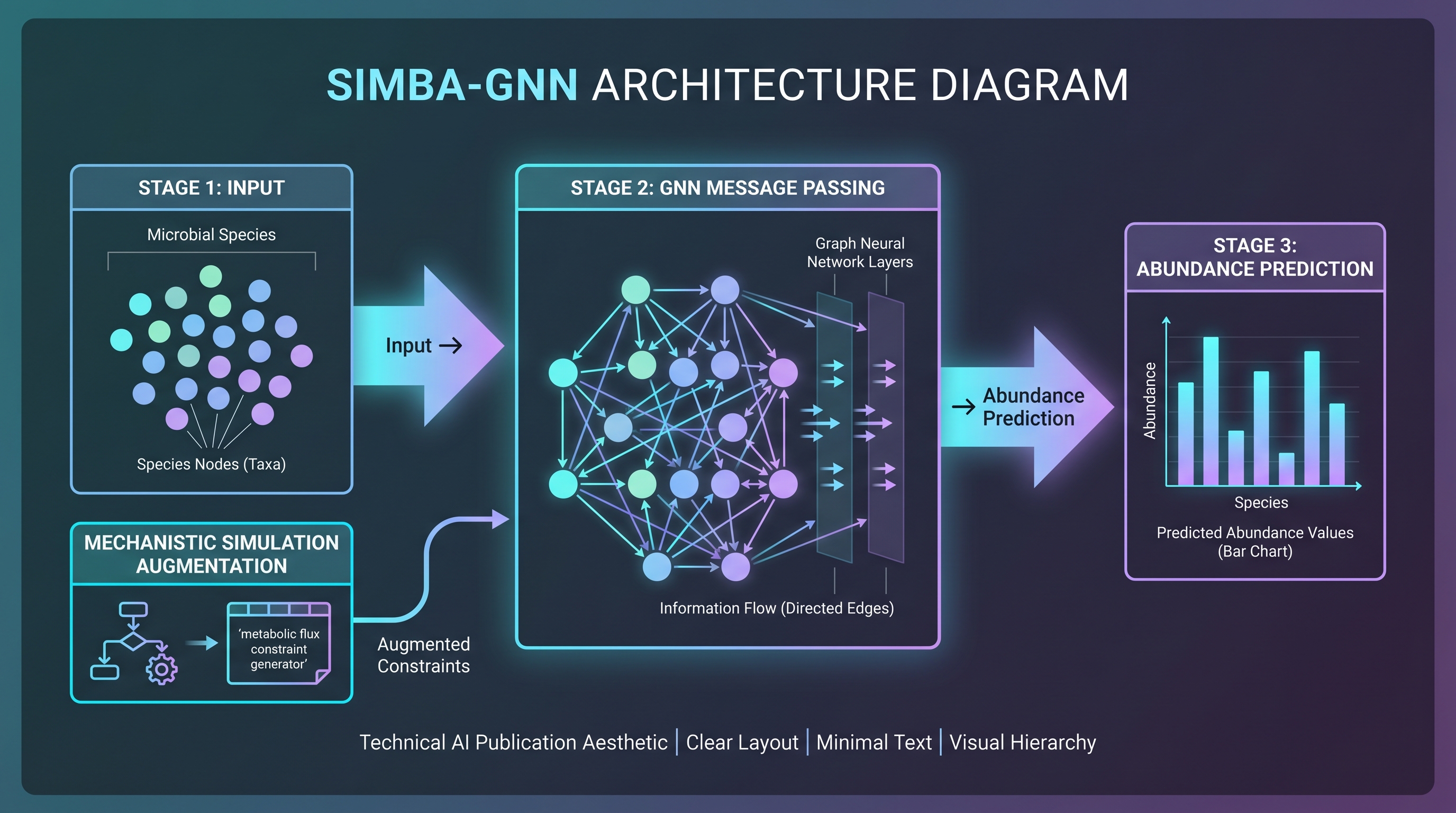
The most natural tool for learning microbial communities as graphs is the GNN. Node embeddings absorb information from neighbors via message passing; the resulting community-level representation feeds downstream classifiers or regressors.
SIMBA-GNN [24], published in npj Systems Biology and Applications, is a 2025 product of this thread. SIMBA stands for Simulation-augmented Microbiome Abundance GNN. It addresses a known weakness of purely data-trained GNNs — poor generalization out of distribution, e.g. to a new perturbation such as applying a novel active — by augmenting the training signal with ODE-based mechanistic simulator output. Inputs are 16S- or shotgun-derived relative-abundance vectors; outputs are abundance predictions at arbitrary forward time. Validation has been mostly gut-cohort-based; the authors explicitly note skin-cohort extension is feasible without changing the domain framework. A skin-cohort readout has not yet appeared.
Earlier threads bringing GNNs to microbiome split into two strands. First, DeepMicro-/MetaPhlAn-style embedding learners — classifiers predicting phenotype from taxonomic profile. Second, community-level metabolite-production prediction — predicting which metabolites a community produces via message passing on the species graph. [12]'s meta-review of 200+ AI-microbiome papers documents the rapid adoption of GNNs and Transformers in microbiome AI, with the caveat that "skin coverage is limited compared to gut." [25] phrases the same point more bluntly — "gut-only focus; skin equivalents lag."
A signal from an adjacent domain is worth borrowing. [14] in Science Advances used a cell-cell communication network with interpretable ML to predict cancer-patient response to immune checkpoint inhibitors — a tumor-immunology problem, not microbiome, but the skeleton ("message passing between nodes + interpretability") transfers cleanly. One plausible next move for cosmetic-microbiome modeling is precisely this shape — learn a microbe–host-cell cross-talk network with a GNN, then surface which nodes drive a phenotype via SHAP or attention analysis.
The industrial value of GNNs comes down to compressing the predict–validate loop. Instead of running every wet-lab assay on 10,000 strain-combination synergies, use a GNN to rank the top 50, then validate just those. The logic mirrors the endpoint-specific ML triage of (Chapter 4) — but the input is a community, not a molecule.
6.3 Mechanistic models — what differential equations still answer
If GNNs learn edges from data, mechanistic models assume edges. The oldest tool here is the generalized Lotka–Volterra (gLV) model — an ODE system describing how species abundances evolve given an interaction-coefficient matrix. Direct readouts in cosmetics are sparse, but gLV is standard in gut microbiome work ([12], [25]), and SIMBA-GNN's mechanistic augmentation step itself runs a gLV-like simulator under the hood.
Agent-based skin models operate at a different abstraction level. Each microbe is treated as an agent on a lattice; local concentrations of nutrient, oxygen, pH, and antimicrobials are solved as ODEs while agents grow, move, and die. [28], a mini-review in Biotechnology Journal on organoid-based skin and lung biofilm models, shows that the in vitro counterpart of these agent-based simulations — multi-microbe biofilms grown on 3D skin organoids that produce data under controlled conditions — has rapidly standardized over 2024–2025. [18], published in Nature Communications, is more ambitious: an in vitro micro-ecology platform that reproduces multi-microbe human skin interactions while allowing experimental control, providing ground truth against which mechanistic parameters can be fit. That makes it quantitatively stronger than GNN-only approaches.
Diffusion ODEs for metabolite spatiotemporal distribution also matter increasingly in cosmetics. Predicting how an active ingredient distributes across the stratum corneum over time, and how local concentration impacts microbial communities, requires solving Fickian diffusion coupled with microbial uptake terms in a PDE. This modeling becomes a key input for the AI formulation work of (Chapter 8).
The major strength of mechanistic models is extrapolation. Given perturbations outside the training distribution, a correct mechanism gives reasonable predictions. The major weakness is parameter determination. The gLV interaction-coefficient matrix scales as the square of species count, and measuring every pair experimentally for a skin community is impractical. That is why hybrids like SIMBA-GNN are natural — learn coefficients from data while regularizing them with mechanism.
6.4 Multi-omics integration — 16S + metabolome + transcriptome
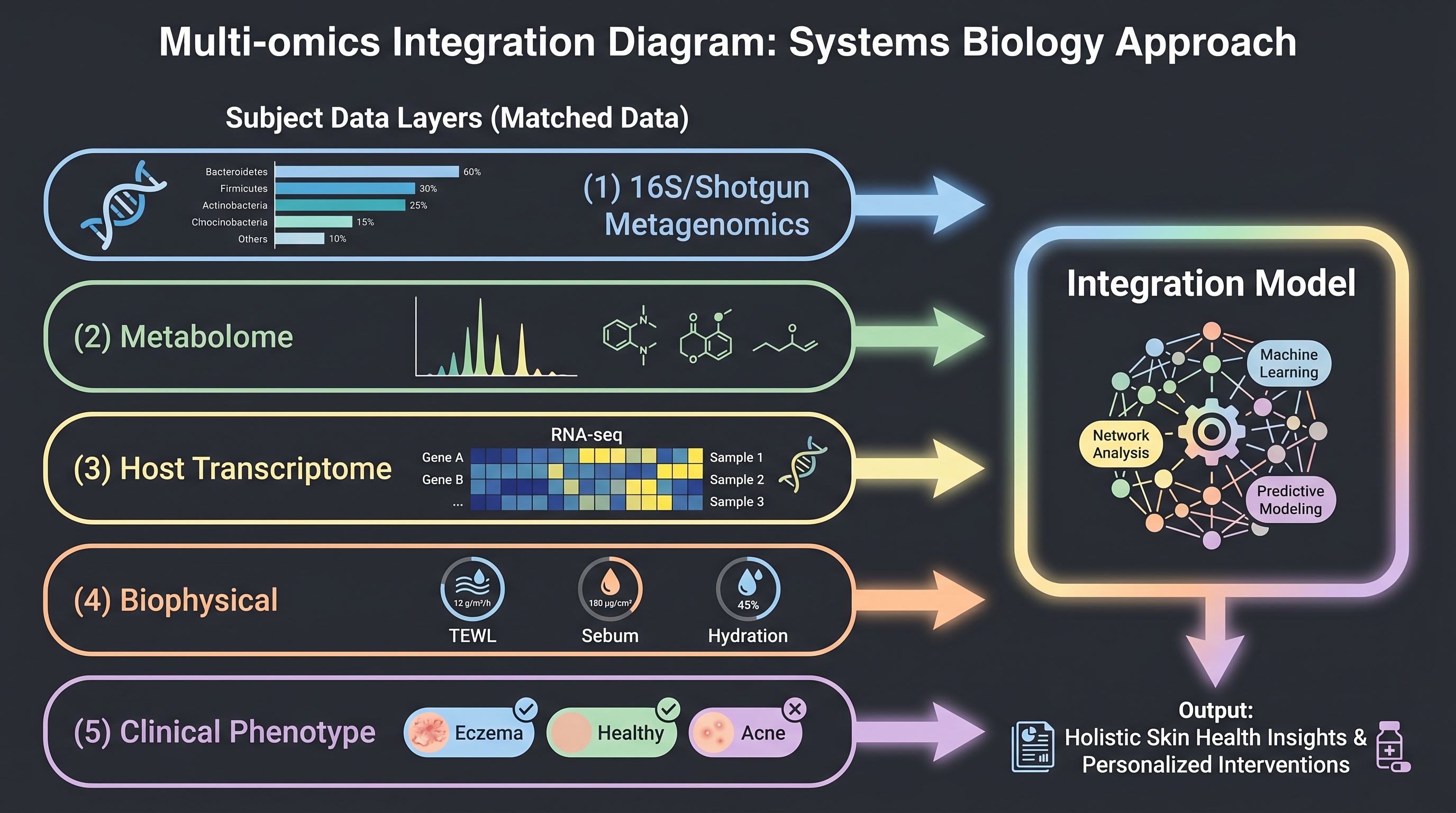
Community modeling rarely has a single input. Models become most powerful when 16S (or shotgun) + metabolome + skin transcriptome are matched at the same time point in the same individual. Reference cohorts with that matching are surprisingly few.
- HMP / iHSMGC — taxonomic profile is curated, but metabolome and transcriptome are absent. iHSMGC [15] catalogs 10.94M genes, of which ~45% are invisible to HMP — a quantitative anchor of how population-locked reference catalogs can be.
- Unilever 30K — disclosed in [26]. 30,000 samples and 5 billion data points are described as matching 16S + biophysical + (some) metabolomics. No external access.
- COSMAX × Dankook FACE-LINK — 950 Korean subjects, published in Frontiers in Cellular and Infection Microbiology by [19]. Matches 16S + biophysical (hydration, TEWL, melanin, erythema). The largest matched cohort released externally by Korean industry.
- L'Oréal internal — Modjoul / Modiface device data plus the ~10,000-strain collection absorbed in the 2023 Lactobio acquisition. Multi-omics integration level is undisclosed.
- iHSMGC + ZOE PREDICT — not directly matched, but [2] PREDICT 1 (1,098 subjects, gut microbiome + diet + metabolic phenotype) functions as the reference for skin-gut axis modeling in (Chapter 11).
This data landscape is what sequencing depth ultimately means — the trade-offs among 16S, shotgun, and long-read described in (Chapter 3) map directly to model input dimensions. With 16S alone you have species-level abundance vectors; shotgun adds functional-gene (KO/EggNOG) features; metabolomics adds a metabolite vector; transcriptomics adds host response — each step increases the causal hypotheses the model can capture. But each step also shrinks the public data available. The microbiome-ML failure modes that [22] enumerated (compositionality, high dimensionality, batch effect, leakage) all worsen at multi-omics integration: each omics layer brings its own batch effect.
In this context, [4], published in Nature, has outsized importance — the finding that skin produces antibodies autonomously and that those antibodies regulate microbiome composition adds a new modeling input dimension: a skin-immune signature. Expect to see it return as an input to the clinical simulators of (Chapter 9).
6.5 The digital twin — who is calling what a twin
"Digital twin," "skin twin," and "microbiome twin" have proliferated in industry vocabulary since 2024. The trouble is that the definition shifts each time. This chapter names the definition gap.
Technical definition (the canonical one). A digital twin (1) represents the current state of a real system as data, (2) predicts future states under perturbations applied to that system, and (3) updates itself by comparing predictions against measurements — a living model. This definition is stable in manufacturing, aviation, and urban-infrastructure literature.
Industry usage (cosmetics / microbiome). Often borrows only some of the above.
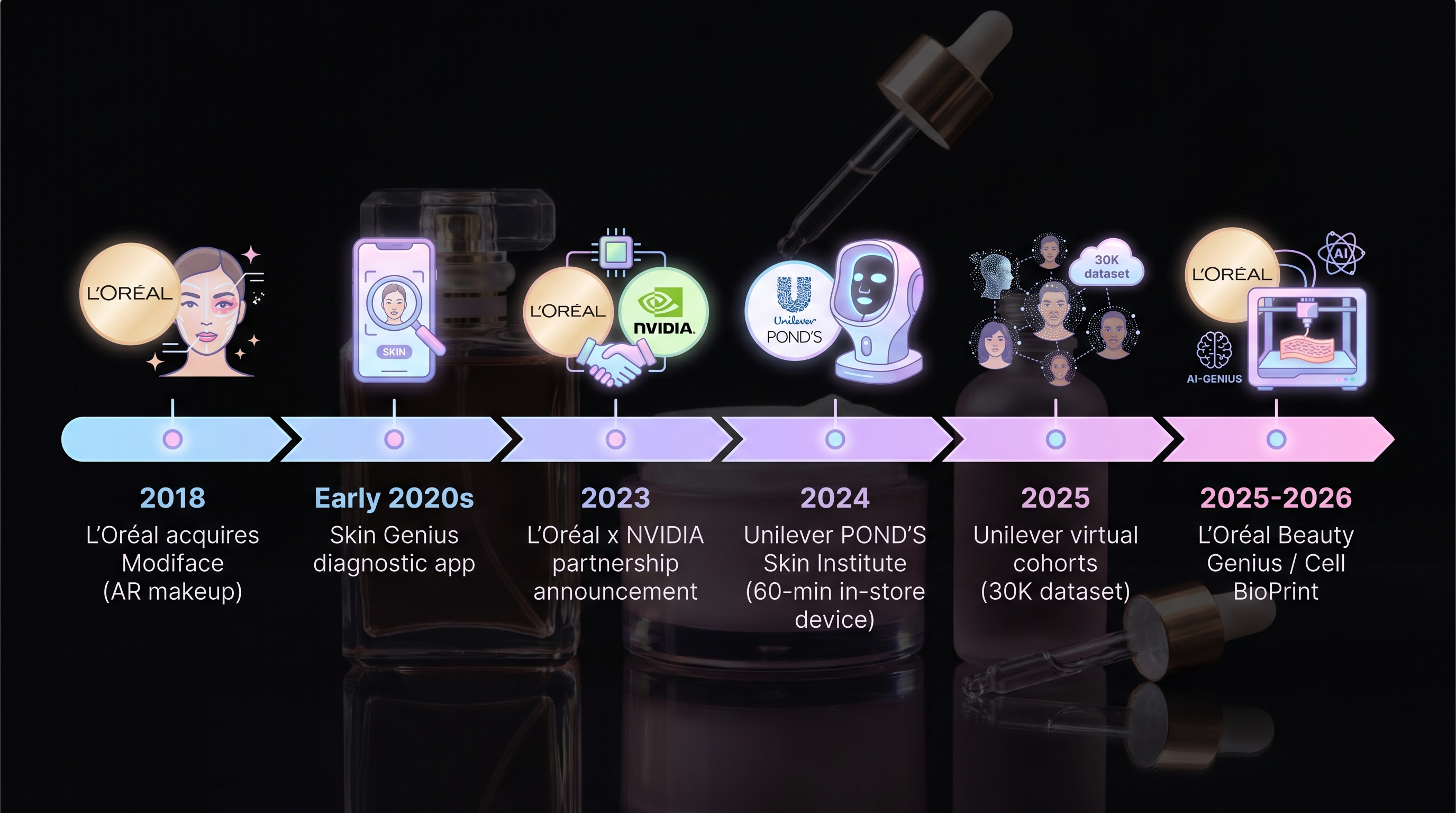
- L'Oréal Skin Genius / Beauty Genius / Cell BioPrint — [17] disclosed AI diagnostic tools that produce skin-state assessments and product recommendations from images and questionnaires. No external validation readout exists. [7]'s PRISMA review explicitly notes that "digital-twin claims rest on commercial demos without peer-reviewed validation."
- Unilever virtual cohorts — [26] disclosed at SXSW a 2,500-subject simulated cohort, reporting 60% faster consumer insight, formulation cycles shrinking from 5–6 to 1–2, and 75% faster claims development. Operational KPIs are reported; generalization error is not ([27]).
- POND's Skin Institute analyzer — [23] launched a 60-minute in-store device with microbiome swab → product recommendation workflow. Performance as a real-time decision tool is undisclosed.
- COSMAX × Bertis skin proteome — [5] disclosed a proteome-based aging model from Korean industry. The "twin" word is not used explicitly, but the structure (deriving recommendations from individual-level proteomic signatures) is from the same lineage.
- Infant microbiome digital twin — [9]: a U. Chicago group's PNAS / npj Biofilms paper. It is an infant gut twin, not skin, but it is the cleanest peer-reviewed instantiation of the term in the corpus — input, prediction, ground truth, and generalization are all explicitly defined.
One pattern emerges from this list — the cosmetic industry's "digital twin" is a marketing twin more than a methodological twin. Just as [7], in their PRISMA-grade review of 74 papers, finds zero externally peer-reviewed clinical readouts for AI-designed cosmetic actives, the same review finds essentially no externally validated digital twins. The most honest single line of this chapter is this: most of what industry currently calls a "twin" is a recommendation system with a personalization layer on top. That is not without value — but calling it a twin that predicts and validates microbiome dynamics requires more than what is currently published.
6.6 The foundation-model gap — addressing Gap 2 head-on
The critical-analyst's heaviest message for this chapter is this. As of May 2026, no skin-microbiome foundation model exists at ESM3 scale.
The comparison anchor is clear. On the protein side, ESM3 [8] is a 98B-parameter model trained on the equivalent of 500 million years of evolution, with a 7B variant released openly. Above it, AlphaFold 2/3 [10] made PDB, UniProt, and AlphaFold DB into reference infrastructure. If the protein inflection was public assets + foundation model, the microbiome side is missing both.
Candidates on the microbiome side. [12]'s 200+ paper synthesis records rapidly rising Transformer/GNN/generative adoption in microbiome AI, but skin coverage is "limited compared to gut." [6] names COMEBin as 2024 SoTA, but that is a metagenomic binner, not a foundation model. [25] flatly states "gut-only focus; skin equivalents lag."
Why the gap. Four reasons.
- Training-corpus privacy — the data scale needed for foundation-model training (tens to hundreds of thousands of individuals with matched multi-omics) sits inside industry. Unilever 30K, L'Oréal internal, COSMAX FACE-LINK 950, BGI/iHSMGC — the largest assets are all private ([15], [19], [26]).
- Fragmentation of consent and ethics — skin-microbiome sampling requires IRB approval and consent scope varies per study. Cross-cohort merging for foundation-model training faces legal barriers.
- Multimodality — proteins admit a relatively unified representation (sequence and structure); the microbiome involves 16S, shotgun, metabolome, transcriptome, biophysical, and clinical modalities, all different. Defining a unified representation is itself an unsolved problem.
- The time dimension — microbiomes are dynamical systems that change with time, not equilibrium structures like proteins. There is no consensus on what timestep representation a foundation model should learn.
What would a foundation model unlock if it existed. At least three things: (a) zero-shot community-dynamics prediction — predicting community shift under an unseen perturbation (a new active ingredient), (b) cross-cohort generalization — a Korean-trained model extrapolating safely to EU and US cohorts, (c) multi-omics stitching — imputing missing modalities for new individuals who only have some modalities measured. None are currently feasible.
The two plausible solution directions. Data cooperatives — industry + academia using federated learning or differential privacy to train a shared model without sharing raw data. Just as BGI led the iHSMGC project, Korean / Chinese / EU industry could form a cooperative. Expanding public reference cohorts — successor cohorts to HMP / iHSMGC that match multi-omics and are released openly. (Chapter 12) analyzes both possibilities at the blueprint level.
This gap is the ceiling of cosmetic-microbiome AI. Until it closes, every task-specific model is bound by its training-corpus size. That is the most direct meaning of Gap 2.
6.7 Limits and risks
The promise of community modeling and twins has a matching set of risks, and they need to be named.
Overfit and out-of-distribution failure — Unilever's 2,500-subject virtual cohort has no externally validated description of how the simulated population was sampled. The pattern of a model scoring 5/5 on the training cohort and 2/5 on a new population — well documented in radiology and drug-efficacy prediction — is not unlikely for microbiome twins. [7] explicitly cites "limited geographic diversity, darker phototypes underrepresented" as a PRISMA-level limitation, and ethnicity / age / environment biases directly impact twin generalization.
The marketing-vs-science gap — the distance between industry KPIs (60% faster, 75% faster) and peer-reviewed validation tracks with Gap 15 from (Chapter 4). When twin claims become marketing weapons, consumers and R&D decision-makers lack the vocabulary to separate what has been validated from what is simulated. One direct contribution of this book is to supply that vocabulary.
Privacy and personalization — the microbiome is partially a fingerprint ([20]). The more precisely a twin operates at individual level, the higher the risk of personal identification and re-identification. As cosmetic companies run twins in production, friction against GDPR and Korea's PIPA grows clear — use-case-specific consent scope, and the risk of identity inversion from model weights, both become operational issues.
Model drift — microbiomes shift slowly with season, diet, climate, and age ([21] reported 1–2 year stability, site-dependent). A twin trained in 2024 making decisions in 2027 accumulates drift as forecast error. Continual learning and re-training cadences must become part of twin operations — this is a system-operations problem, not just a deployment problem.
These limits are not model-engineering problems. They are data, governance, and validation problems. Changing the model architecture will not close them. (Chapter 12) takes up that governance blueprint substantively.
6.8 Open Questions
- Minimum data scale for a foundation model — how many individuals with matched multi-omics are needed before a skin-microbiome foundation model shows meaningful zero-shot generalization? Even an order-of-magnitude estimate is hard relative to proteins (UniProt 250M+). If the learning curve flattens at 50K, an industry cooperative is feasible. If 1M+ is needed, no cooperative can reach there.
- Twin validation criteria — what minimum conditions justify calling a model a "twin"? (a) prospective prediction, (b) ground-truth measurement, (c) error quantification, (d) generalization to an unseen population — these four define [9], but there is no consensus on how to translate them to cosmetic endpoints (e.g. "hydration score at week 8").
- Long-term stability — how different is a user's microbiome baseline 5 years after a twin is trained? The 1–2 year stability data from [21] may not cover cosmetic product life cycles (typically 5 years) — and there is no industry standard for drift monitoring.
- Individual twin vs cohort twin — "skin twin" is often ambiguous between an individual twin and a population-segment twin. Unilever's 2,500-subject virtual cohort is closer to a segment twin; L'Oréal's Skin Genius asserts an individual twin. The two approaches differ in data, validation, and legal liability — and the industry distinction has not clarified.
- Synthetic data's role — how well do virtual-cohort simulated samples represent the real distribution? Mechanistic-ODE-based augmentation ([24]) is valuable as a training signal, but using it as validation data is a separate risk. The line between training and validation is often blurred in industry KPI disclosures.
References
- Abramson, J., Adler, J., Dunger, J. et al. (2024). Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500.
- Asnicar, F., Berry, S. E., Valdes, A. M. et al. (2021). Microbiome connections with host metabolism and habitual diet from 1,098 deeply phenotyped individuals. Nature Medicine 27, 321–332.
- Belkaid, Y. and Segre, J. A. (2014). Dialogue between skin microbiota and immunity. Science 346(6212), 954–959. [Belkaid et al., 2014]
- Belkaid, Y. et al. (2024). Skin autonomous antibody production regulates host-microbiota interactions. Nature, December 2024. [Belkaid et al., 2024]
- COSMAX × Bertis (2024). COSMAX × Bertis skin proteome anti-aging partnership. The Monodist, 2024–2025. [COSMAX × Bertis, 2024]
- Deep learning microbiome review — Wang, T., Yang, L. et al. (2024). Deep learning in microbiome analysis: a comprehensive review of neural network models. Frontiers in Microbiology 15:1516667. [Deep learning microbiome review, 2024]
- Haykal, D., Flament, F., Amar, D. et al. (2025). Cosmetogenomics unveiled: a systematic review of AI, genomics, and the future of personalized skincare. Frontiers in Artificial Intelligence 8:1660356.
- Hayes, T., Rao, R., Akin, H. et al. (2025). Simulating 500 million years of evolution with a language model (ESM3). Science, 2025.
- Infant microbiome digital twin authors — U. Chicago group (2024). A digital twin of the infant microbiome to predict neurodevelopmental deficits. PNAS / npj Biofilms, 2024. [Infant microbiome digital twin, 2024]
- Jumper, J., Evans, R., Pritzel, A. et al. (2021). Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589.
- Khadka, V. D. et al. (2024). Commensal skin bacteria exacerbate inflammation on damaged skin barriers. Journal of Investigative Dermatology, 2024. [Khadka et al., 2024]
- Wang, X.-W., Wang, T., Liu, Y.-Y. (2024). Artificial Intelligence for Microbiology and Microbiome Research. arXiv preprint 2411.01098. [Wang et al., 2024]
- Krismer, B., Peschel, A. et al. (2024). Commensal production of broad-spectrum antimicrobial peptide polyene eliminates nasal S. aureus (epifadin). Nature Microbiology, 2024.
- Lee, J., Kim, D., Kong, J. et al. (2024). Cell-cell communication network-based interpretable machine learning predicts cancer patient response to immune checkpoint inhibitors. Science Advances 10:eadj0785.
- Li, Z., Xia, J., Jiang, L. et al. (2021). Characterization of the human skin resistome and identification of two microbiota cutotypes (iHSMGC catalog). Microbiome 9:47.
- Li, Z., Xia, J., Wang, J. (2025). Unveiling strain-level dynamics in the human skin microbiome (Preview of Jacob et al.)00097-6). Cell Host & Microbe 33(5):615–617.
- L'Oréal R&I Press (2024). L'Oréal Beauty Tech leadership at VivaTech 2024 — AI Skin Genius, Modiface, microbiome direction. L'Oréal press, May 2024. [L'Oréal VivaTech, 2024]
- Microecology in vitro authors (2025). Microecology in vitro model replicates the human skin microbiome interactions. Nature Communications, 2025. [Microecology in vitro, 2025]
- Mun, S., Jo, H., Heo, Y. M. et al. (2025). Skin microbiome-biophysical association: a first integrative approach to classifying Korean skin types and aging groups (FACE-LINK). Frontiers in Cellular and Infection Microbiology 15:1561590.
- Oh, J., Byrd, A. L., Deming, C. et al. (2014). Biogeography and individuality shape function in the human skin metagenome. Nature 514, 59–64. [Oh et al., 2014]
- Oh, J., Byrd, A. L., Park, M. et al. (2016). Temporal Stability of the Human Skin Microbiome30399-3). Cell 165(4), 854–866. [Oh et al., 2016]
- Papoutsoglou, G., Tarazona, S., Lopes, M. B. et al. (2023). Machine learning approaches in microbiome research: challenges and best practices. Frontiers in Microbiology 14:1261889.
- POND's / Unilever (2024). POND's Skin Institute microbiome analyzer — 60-minute in-store consumer device. Unilever press, May 2024. [POND's, 2024]
- SIMBA-GNN authors (2025). SIMBA-GNN: Simulation-augmented Microbiome Abundance GNN. npj Systems Biology and Applications, 2025. [SIMBA-GNN, 2025]
- Transformer-microbiome authors (2024). Transformer Models, Graph Networks, and Generative AI in Gut Microbiome Research. Bioengineering 13(2):144. [Transformermicrobiome, 2024]
- Unilever Beauty & Wellbeing R&D (2025). How Unilever's pioneering skin microbiome research is shaping product innovation. Unilever news; SXSW 2025 + R&D page coverage. [Unilever, 2025]
- Unilever (2026). Unilever 2026 forward outlook — AI transforming Beauty & Wellbeing innovation. Unilever news, 2026. [Unilever, 2026]
- Verma, P. et al. (2025). Organoid-Based Skin and Lung Biofilm Models — Mini Review. Biotechnology Journal, 2025. [Verma et al., 2025]